!pip install matplotlib s3fs "xarray[io]"
Show code cell output
Hide code cell output
Requirement already satisfied: matplotlib in /usr/lib/python3/dist-packages (3.1.2)
Requirement already satisfied: s3fs in /home/deploy/.local/lib/python3.8/site-packages (2024.10.0)
Requirement already satisfied: xarray[io] in /home/deploy/.local/lib/python3.8/site-packages (2023.1.0)
Requirement already satisfied: aiohttp!=4.0.0a0,!=4.0.0a1 in /home/deploy/.local/lib/python3.8/site-packages (from s3fs) (3.10.11)
Requirement already satisfied: fsspec==2024.10.0.* in /home/deploy/.local/lib/python3.8/site-packages (from s3fs) (2024.10.0)
Requirement already satisfied: aiobotocore<3.0.0,>=2.5.4 in /home/deploy/.local/lib/python3.8/site-packages (from s3fs) (2.22.0)
Requirement already satisfied: pandas>=1.3 in /home/deploy/.local/lib/python3.8/site-packages (from xarray[io]) (2.0.3)
Requirement already satisfied: packaging>=21.3 in /usr/local/lib/python3.8/dist-packages (from xarray[io]) (24.1)
Requirement already satisfied: numpy>=1.20 in /home/deploy/.local/lib/python3.8/site-packages (from xarray[io]) (1.24.4)
Requirement already satisfied: scipy; extra == "io" in /home/deploy/.local/lib/python3.8/site-packages (from xarray[io]) (1.10.1)
Requirement already satisfied: zarr; extra == "io" in /home/deploy/.local/lib/python3.8/site-packages (from xarray[io]) (2.16.1)
Requirement already satisfied: cfgrib; extra == "io" in /home/deploy/.local/lib/python3.8/site-packages (from xarray[io]) (0.9.15.0)
Requirement already satisfied: h5netcdf; extra == "io" in /home/deploy/.local/lib/python3.8/site-packages (from xarray[io]) (1.1.0)
Requirement already satisfied: rasterio; extra == "io" in /home/deploy/.local/lib/python3.8/site-packages (from xarray[io]) (1.3.11)
Requirement already satisfied: netCDF4; extra == "io" in /home/deploy/.local/lib/python3.8/site-packages (from xarray[io]) (1.7.2)
Requirement already satisfied: cftime; extra == "io" in /home/deploy/.local/lib/python3.8/site-packages (from xarray[io]) (1.6.4.post1)
Requirement already satisfied: pooch; extra == "io" in /home/deploy/.local/lib/python3.8/site-packages (from xarray[io]) (1.8.2)
Requirement already satisfied: pydap; python_version < "3.10" and extra == "io" in /home/deploy/.local/lib/python3.8/site-packages (from xarray[io]) (3.4.1)
Requirement already satisfied: multidict<7.0,>=4.5 in /home/deploy/.local/lib/python3.8/site-packages (from aiohttp!=4.0.0a0,!=4.0.0a1->s3fs) (6.1.0)
Requirement already satisfied: aiosignal>=1.1.2 in /home/deploy/.local/lib/python3.8/site-packages (from aiohttp!=4.0.0a0,!=4.0.0a1->s3fs) (1.3.1)
Requirement already satisfied: yarl<2.0,>=1.12.0 in /home/deploy/.local/lib/python3.8/site-packages (from aiohttp!=4.0.0a0,!=4.0.0a1->s3fs) (1.15.2)
Requirement already satisfied: frozenlist>=1.1.1 in /home/deploy/.local/lib/python3.8/site-packages (from aiohttp!=4.0.0a0,!=4.0.0a1->s3fs) (1.5.0)
Requirement already satisfied: aiohappyeyeballs>=2.3.0 in /home/deploy/.local/lib/python3.8/site-packages (from aiohttp!=4.0.0a0,!=4.0.0a1->s3fs) (2.4.4)
Requirement already satisfied: async-timeout<6.0,>=4.0; python_version < "3.11" in /home/deploy/.local/lib/python3.8/site-packages (from aiohttp!=4.0.0a0,!=4.0.0a1->s3fs) (5.0.1)
Requirement already satisfied: attrs>=17.3.0 in /usr/lib/python3/dist-packages (from aiohttp!=4.0.0a0,!=4.0.0a1->s3fs) (19.3.0)
Requirement already satisfied: wrapt<2.0.0,>=1.10.10 in /home/deploy/.local/lib/python3.8/site-packages (from aiobotocore<3.0.0,>=2.5.4->s3fs) (1.17.2)
Requirement already satisfied: botocore<1.37.4,>=1.37.2 in /home/deploy/.local/lib/python3.8/site-packages (from aiobotocore<3.0.0,>=2.5.4->s3fs) (1.37.3)
Requirement already satisfied: jmespath<2.0.0,>=0.7.1 in /usr/lib/python3/dist-packages (from aiobotocore<3.0.0,>=2.5.4->s3fs) (0.9.4)
Requirement already satisfied: python-dateutil<3.0.0,>=2.1 in /home/deploy/.local/lib/python3.8/site-packages (from aiobotocore<3.0.0,>=2.5.4->s3fs) (2.9.0.post0)
Requirement already satisfied: aioitertools<1.0.0,>=0.5.1 in /home/deploy/.local/lib/python3.8/site-packages (from aiobotocore<3.0.0,>=2.5.4->s3fs) (0.12.0)
Requirement already satisfied: tzdata>=2022.1 in /home/deploy/.local/lib/python3.8/site-packages (from pandas>=1.3->xarray[io]) (2025.2)
Requirement already satisfied: pytz>=2020.1 in /home/deploy/.local/lib/python3.8/site-packages (from pandas>=1.3->xarray[io]) (2025.2)
Requirement already satisfied: asciitree in /home/deploy/.local/lib/python3.8/site-packages (from zarr; extra == "io"->xarray[io]) (0.3.3)
Requirement already satisfied: fasteners in /home/deploy/.local/lib/python3.8/site-packages (from zarr; extra == "io"->xarray[io]) (0.19)
Requirement already satisfied: numcodecs>=0.10.0 in /home/deploy/.local/lib/python3.8/site-packages (from zarr; extra == "io"->xarray[io]) (0.12.1)
Requirement already satisfied: click in /usr/lib/python3/dist-packages (from cfgrib; extra == "io"->xarray[io]) (7.0)
Requirement already satisfied: eccodes>=0.9.8 in /home/deploy/.local/lib/python3.8/site-packages (from cfgrib; extra == "io"->xarray[io]) (2.42.0)
Requirement already satisfied: h5py in /home/deploy/.local/lib/python3.8/site-packages (from h5netcdf; extra == "io"->xarray[io]) (3.11.0)
Requirement already satisfied: affine in /home/deploy/.local/lib/python3.8/site-packages (from rasterio; extra == "io"->xarray[io]) (2.4.0)
Requirement already satisfied: snuggs>=1.4.1 in /home/deploy/.local/lib/python3.8/site-packages (from rasterio; extra == "io"->xarray[io]) (1.4.7)
Requirement already satisfied: setuptools in /usr/lib/python3/dist-packages (from rasterio; extra == "io"->xarray[io]) (45.2.0)
Requirement already satisfied: certifi in /usr/lib/python3/dist-packages (from rasterio; extra == "io"->xarray[io]) (2019.11.28)
Requirement already satisfied: cligj>=0.5 in /home/deploy/.local/lib/python3.8/site-packages (from rasterio; extra == "io"->xarray[io]) (0.7.2)
Requirement already satisfied: click-plugins in /home/deploy/.local/lib/python3.8/site-packages (from rasterio; extra == "io"->xarray[io]) (1.1.1.2)
Requirement already satisfied: importlib-metadata; python_version < "3.10" in /usr/lib/python3/dist-packages (from rasterio; extra == "io"->xarray[io]) (1.5.0)
Requirement already satisfied: requests>=2.19.0 in /usr/lib/python3/dist-packages (from pooch; extra == "io"->xarray[io]) (2.22.0)
Requirement already satisfied: platformdirs>=2.5.0 in /home/deploy/.local/lib/python3.8/site-packages (from pooch; extra == "io"->xarray[io]) (4.3.6)
Requirement already satisfied: six>=1.4.0 in /usr/lib/python3/dist-packages (from pydap; python_version < "3.10" and extra == "io"->xarray[io]) (1.14.0)
Requirement already satisfied: beautifulsoup4 in /home/deploy/.local/lib/python3.8/site-packages (from pydap; python_version < "3.10" and extra == "io"->xarray[io]) (4.13.4)
Requirement already satisfied: Webob in /home/deploy/.local/lib/python3.8/site-packages (from pydap; python_version < "3.10" and extra == "io"->xarray[io]) (1.8.9)
Requirement already satisfied: docopt in /usr/lib/python3/dist-packages (from pydap; python_version < "3.10" and extra == "io"->xarray[io]) (0.6.2)
Requirement already satisfied: Jinja2 in /usr/local/lib/python3.8/dist-packages (from pydap; python_version < "3.10" and extra == "io"->xarray[io]) (3.1.4)
Requirement already satisfied: typing-extensions>=4.1.0; python_version < "3.11" in /home/deploy/.local/lib/python3.8/site-packages (from multidict<7.0,>=4.5->aiohttp!=4.0.0a0,!=4.0.0a1->s3fs) (4.13.2)
Requirement already satisfied: idna>=2.0 in /usr/lib/python3/dist-packages (from yarl<2.0,>=1.12.0->aiohttp!=4.0.0a0,!=4.0.0a1->s3fs) (2.8)
Requirement already satisfied: propcache>=0.2.0 in /home/deploy/.local/lib/python3.8/site-packages (from yarl<2.0,>=1.12.0->aiohttp!=4.0.0a0,!=4.0.0a1->s3fs) (0.2.0)
Requirement already satisfied: urllib3<1.27,>=1.25.4; python_version < "3.10" in /usr/lib/python3/dist-packages (from botocore<1.37.4,>=1.37.2->aiobotocore<3.0.0,>=2.5.4->s3fs) (1.25.8)
Requirement already satisfied: cffi in /home/deploy/.local/lib/python3.8/site-packages (from eccodes>=0.9.8->cfgrib; extra == "io"->xarray[io]) (1.17.1)
Requirement already satisfied: findlibs in /home/deploy/.local/lib/python3.8/site-packages (from eccodes>=0.9.8->cfgrib; extra == "io"->xarray[io]) (0.1.1)
Requirement already satisfied: pyparsing>=2.1.6 in /usr/lib/python3/dist-packages (from snuggs>=1.4.1->rasterio; extra == "io"->xarray[io]) (2.4.6)
Requirement already satisfied: soupsieve>1.2 in /home/deploy/.local/lib/python3.8/site-packages (from beautifulsoup4->pydap; python_version < "3.10" and extra == "io"->xarray[io]) (2.7)
Requirement already satisfied: MarkupSafe>=2.0 in /usr/local/lib/python3.8/dist-packages (from Jinja2->pydap; python_version < "3.10" and extra == "io"->xarray[io]) (2.1.5)
Requirement already satisfied: pycparser in /home/deploy/.local/lib/python3.8/site-packages (from cffi->eccodes>=0.9.8->cfgrib; extra == "io"->xarray[io]) (2.22)
import zarr
import numpy as np
import xarray as xr
import matplotlib.pyplot as plt
from matplotlib.colors import LogNorm
from scipy.signal import stft
Level 2 Data#
In this notebook we demonstrate the variety of different data available in the FAIR MAST dataset. In this example we are using level 2 MAST data, which includes cropping, interpolation, calibration, etc. of each signal, as well as mapping each diagnostic group try and follow IMAS naming convetions. The level 2 data is well-indexed data and follows the FAIR principles. Shots are also filtered using the plasma current to remove shots which were only used for testing, comissioning, machine calibration etc.
First we need to connect to the remote S3 storage bucket to access the data. Each shot from MAST is stored as a seperate Zarr file.
Using fsspec and xarray we can remotely read data directly over the web. In the example below we also turn on local file caching, allowing us to avoid reading over the network multiple times.
shot_id = 30421
endpoint_url = 'https://s3.echo.stfc.ac.uk'
url = f's3://mast/level2/shots/{shot_id}.zarr'
# Get a handle to the remote file
store = zarr.storage.FsspecStore.from_url(
url,
storage_options=dict(
protocol='simplecache',
target_protocol="s3",
cache_storage='.cache',
target_options=dict(
anon=True, endpoint_url=endpoint_url, asynchronous=True
)
)
)
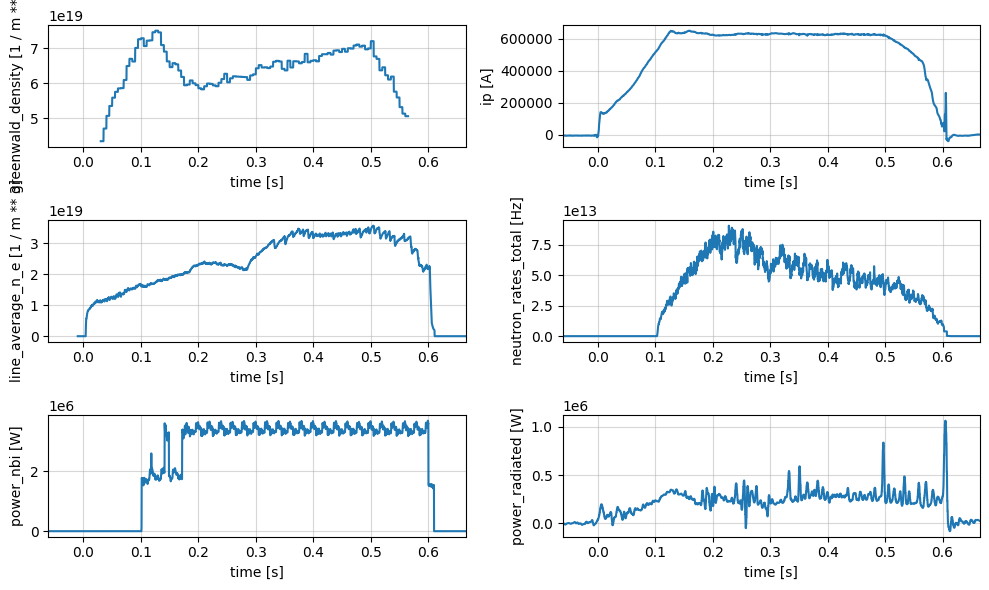
Summary Profiles#
The summary group provides a collection of general physics quantities for an experiment.
Show code cell source
Hide code cell source
def plot_1d_profiles(profiles: xr.Dataset):
"""Helper function for plotting 1D profiles"""
n = int(np.ceil(len(profiles.data_vars) / 2))
fig, axes = plt.subplots(n, 2, figsize=(10, 2*n))
axes = axes.flatten()
for i, name in enumerate(profiles.data_vars.keys()):
profiles[name].plot(x='time', ax=axes[i])
for ax in axes:
ax.grid('on', alpha=0.5)
ax.set_xlim(profiles.time.min(), profiles.time.max())
plt.tight_layout()
profiles = xr.open_zarr(store, group='summary')
plot_1d_profiles(profiles)
profiles
<xarray.Dataset> Size: 163kB
Dimensions: (time: 2906)
Coordinates:
* time (time) float64 23kB -0.0612 -0.06095 ... 0.6648 0.665
Data variables:
greenwald_density (time) float64 23kB ...
ip (time) float64 23kB ...
line_average_n_e (time) float64 23kB ...
neutron_rates_total (time) float64 23kB ...
power_nbi (time) float64 23kB ...
power_radiated (time) float64 23kB ...
Attributes:
name: summary
description: Summary of physics quantities from a simulation or an expe...
imas: summary
license_name: Creative Commons 4.0 BY-SA
license_url: https://creativecommons.org/licenses/by-sa/4.0/- time: 2906
- time(time)float64-0.0612 -0.06095 ... 0.6648 0.665
- units :
- s
array([-0.0612 , -0.06095, -0.0607 , ..., 0.66455, 0.6648 , 0.66505], shape=(2906,))
- greenwald_density(time)float64...
- data_block_size :
- 115724
- description :
- label :
- Greenwald density limit
- name :
- greenwald_density
- uda_name :
- ESM_N_GREENWALD
- units :
- 1 / m ** 3
[2906 values with dtype=float64]
- ip(time)float64...
- data_block_size :
- 474428
- description :
- Total plasma current (toroidal component). Positive sign means anti-clockwise when viewed from above.
- label :
- Plasma Current
- name :
- ip
- uda_name :
- AMC_PLASMA CURRENT
- imas :
- summary.global_quantities.ip
- units :
- A
[2906 values with dtype=float64]
- line_average_n_e(time)float64...
- data_block_size :
- 507644
- description :
- label :
- Ne_bar
- name :
- line_average_n_e
- uda_name :
- ESM_NE_BAR
- imas :
- summary.line_average.n_e.value
- units :
- 1 / m ** 3
[2906 values with dtype=float64]
- neutron_rates_total(time)float64...
- data_block_size :
- 516936
- description :
- Total neutron rate from all reactions
- label :
- neutrons
- name :
- neutron_rates_total
- uda_name :
- ANU_NEUTRONS
- imas :
- summary.fusion.neutron_rates.total
- units :
- Hz
[2906 values with dtype=float64]
- power_nbi(time)float64...
- data_block_size :
- 914444
- description :
- Total NBI power coupled to the plasma
- label :
- P(SS+SW)
- name :
- power_nbi
- uda_name :
- ANB_TOT_SUM_POWER
- imas :
- summary.heating_current_drive.power_nbi
- units :
- W
[2906 values with dtype=float64]
- power_radiated(time)float64...
- data_block_size :
- 204428
- description :
- label :
- P_Rad_Pol
- name :
- power_radiated
- uda_name :
- ABM_PRAD_POL
- imas :
- summary.global_quantities.power_radiated.value
- units :
- W
[2906 values with dtype=float64]
- timePandasIndex
PandasIndex(Index([-0.06120026111602783, -0.06095026111602783, -0.06070026111602783, -0.06045026111602783, -0.06020026111602783, -0.05995026111602783, -0.05970026111602783, -0.05945026111602783, -0.05920026111602783, -0.05895026111602783, ... 0.6627997388839728, 0.6630497388839728, 0.6632997388839728, 0.6635497388839728, 0.6637997388839728, 0.6640497388839728, 0.6642997388839729, 0.6645497388839728, 0.6647997388839728, 0.6650497388839728], dtype='float64', name='time', length=2906))
- name :
- summary
- description :
- Summary of physics quantities from a simulation or an experiment. Dynamic quantities are either taken at given time slices (indicated in the “time” vector) or time-averaged over an interval (in such case the “time_width” of the interval is indicated and the “time” vector represents the end of each time interval).
- imas :
- summary
- license_name :
- Creative Commons 4.0 BY-SA
- license_url :
- https://creativecommons.org/licenses/by-sa/4.0/
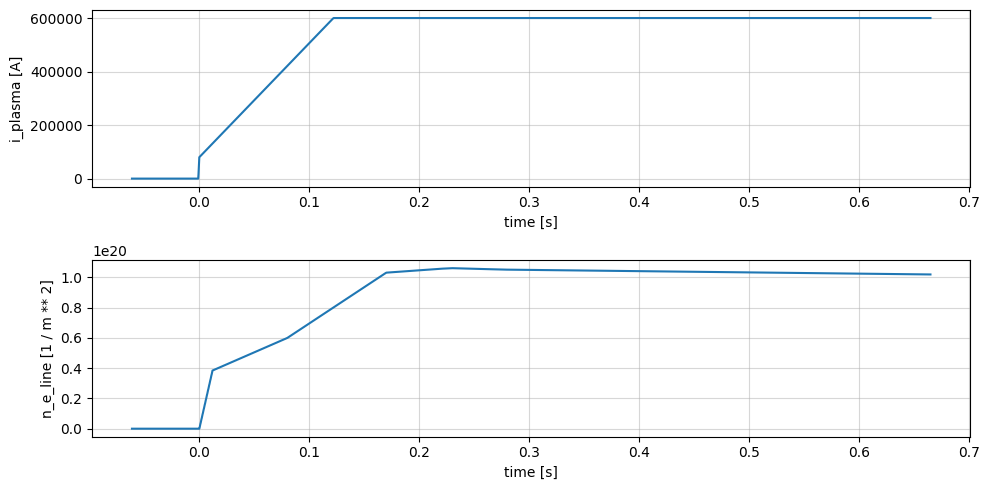
Pulse Schedule#
profiles = xr.open_zarr(store, group='pulse_schedule')
fig, axes = plt.subplots(2, 1, figsize=(10, 5))
axes = axes.flatten()
profiles['i_plasma'].plot(x='time', ax=axes[0])
profiles['n_e_line'].plot(x='time', ax=axes[1])
for ax in axes:
ax.grid('on', alpha=0.5)
plt.tight_layout()
profiles
<xarray.Dataset> Size: 70kB
Dimensions: (time: 2906)
Coordinates:
* time (time) float64 23kB -0.0612 -0.06095 -0.0607 ... 0.6648 0.665
Data variables:
i_plasma (time) float64 23kB ...
n_e_line (time) float64 23kB ...
Attributes:
name: pulse_schedule
description: Description of Pulse Schedule, described by subsystems wav...
imas: pulse_schedule
license_name: Creative Commons 4.0 BY-SA
license_url: https://creativecommons.org/licenses/by-sa/4.0/- time: 2906
- time(time)float64-0.0612 -0.06095 ... 0.6648 0.665
- units :
- s
array([-0.0612 , -0.06095, -0.0607 , ..., 0.66455, 0.6648 , 0.66505], shape=(2906,))
- i_plasma(time)float64...
- data_block_size :
- 450428
- description :
- Plasma current
- label :
- /xdc/ip/t/ipref
- name :
- i_plasma
- uda_name :
- /xdc/ip/t/ipref
- imas :
- pulse_schedule.flux_control.ip
- units :
- A
[2906 values with dtype=float64]
- n_e_line(time)float64...
- data_block_size :
- 450428
- description :
- Line integrated electron density over a line of sight in the whole vacuum chamber
- label :
- /xdc/density/t/nelref
- name :
- n_e_line
- uda_name :
- /xdc/density/t/nelref
- imas :
- pulse_schedule.density_control.n_e_line
- units :
- 1 / m ** 2
[2906 values with dtype=float64]
- timePandasIndex
PandasIndex(Index([-0.06120026111602783, -0.06095026111602783, -0.06070026111602783, -0.06045026111602783, -0.06020026111602783, -0.05995026111602783, -0.05970026111602783, -0.05945026111602783, -0.05920026111602783, -0.05895026111602783, ... 0.6627997388839728, 0.6630497388839728, 0.6632997388839728, 0.6635497388839728, 0.6637997388839728, 0.6640497388839728, 0.6642997388839729, 0.6645497388839728, 0.6647997388839728, 0.6650497388839728], dtype='float64', name='time', length=2906))
- name :
- pulse_schedule
- description :
- Description of Pulse Schedule, described by subsystems waveform references and an envelope around them. The controllers, pulse schedule and SDN are defined in separate IDSs.
- imas :
- pulse_schedule
- license_name :
- Creative Commons 4.0 BY-SA
- license_url :
- https://creativecommons.org/licenses/by-sa/4.0/
Magnetics#
Magnetic diagnostics for equilibrium identification and plasma shape control.
profiles = xr.open_zarr(store, group='magnetics')
fig, axes = plt.subplots(4, 3, figsize=(8, 10))
axes = axes.flatten()
profiles['b_field_pol_probe_ccbv_field'].plot.line(x='time', ax=axes[0], add_legend=False)
profiles['b_field_pol_probe_obv_field'].plot.line(x='time', ax=axes[1], add_legend=False)
profiles['b_field_pol_probe_obr_field'].plot.line(x='time', ax=axes[2], add_legend=False)
profiles['b_field_pol_probe_omv_voltage'].plot.line(x='time_mirnov', ax=axes[3], add_legend=False)
profiles['b_field_pol_probe_cc_field'].plot.line(x='time_mirnov', ax=axes[4], add_legend=False)
profiles['b_field_tor_probe_cc_field'].plot.line(x='time_mirnov', ax=axes[5], add_legend=False)
profiles['b_field_tor_probe_saddle_field'].plot.line(x='time_saddle', ax=axes[6], add_legend=False)
profiles['b_field_tor_probe_saddle_voltage'].plot.line(x='time_saddle', ax=axes[7], add_legend=False)
profiles['b_field_tor_probe_omaha_voltage'].plot.line(x='time_omaha', ax=axes[8], add_legend=False)
profiles['flux_loop_flux'].plot.line(x='time', ax=axes[9], add_legend=False)
profiles['ip'].plot.line(x='time', ax=axes[10], add_legend=False)
for ax in axes:
ax.grid('on', alpha=0.5)
plt.tight_layout()
profiles
<xarray.Dataset> Size: 335MB
Dimensions: (
b_field_pol_probe_cc_channel: 5,
time_mirnov: 363201,
b_field_pol_probe_cc_geometry_channel: 40,
b_field_pol_probe_ccbv_channel: 40,
time: 3633,
...
coordinate: 28,
b_field_tor_probe_saddle_m_geometry_channel: 12,
b_field_tor_probe_saddle_u_geometry_channel: 12,
b_field_tor_probe_saddle_voltage_channel: 12,
flux_loop_channel: 15,
flux_loop_geometry_channel: 44)
Coordinates: (12/25)
* b_field_pol_probe_cc_channel (b_field_pol_probe_cc_channel) <U13 260B ...
* b_field_pol_probe_cc_geometry_channel (b_field_pol_probe_cc_geometry_channel) object 320B ...
* b_field_pol_probe_ccbv_channel (b_field_pol_probe_ccbv_channel) <U10 2kB ...
* b_field_pol_probe_ccbv_geometry_channel (b_field_pol_probe_ccbv_geometry_channel) object 320B ...
* b_field_pol_probe_obr_channel (b_field_pol_probe_obr_channel) <U9 684B ...
* b_field_pol_probe_obr_geometry_channel (b_field_pol_probe_obr_geometry_channel) object 152B ...
... ...
* flux_loop_channel (flux_loop_channel) <U12 720B ...
* flux_loop_geometry_channel (flux_loop_geometry_channel) object 352B ...
* time (time) float64 29kB -0.0612 ...
* time_mirnov (time_mirnov) float64 3MB -0...
* time_omaha (time_omaha) float64 58MB -0...
* time_saddle (time_saddle) float64 291kB ...
Data variables: (12/46)
b_field_pol_probe_cc_field (b_field_pol_probe_cc_channel, time_mirnov) float64 15MB ...
b_field_pol_probe_cc_phi (b_field_pol_probe_cc_geometry_channel) float64 320B ...
b_field_pol_probe_cc_r (b_field_pol_probe_cc_geometry_channel) float64 320B ...
b_field_pol_probe_cc_z (b_field_pol_probe_cc_geometry_channel) float64 320B ...
b_field_pol_probe_ccbv_field (b_field_pol_probe_ccbv_channel, time) float64 1MB ...
b_field_pol_probe_ccbv_length (b_field_pol_probe_ccbv_geometry_channel) float64 320B ...
... ...
b_field_tor_probe_saddle_u_z (b_field_tor_probe_saddle_u_geometry_channel, coordinate) float64 3kB ...
b_field_tor_probe_saddle_voltage (b_field_tor_probe_saddle_voltage_channel, time_saddle) float64 3MB ...
flux_loop_flux (flux_loop_channel, time) float64 436kB ...
flux_loop_r (flux_loop_geometry_channel) float64 352B ...
flux_loop_z (flux_loop_geometry_channel) float64 352B ...
ip (time) float64 29kB -6.039e+...
Attributes:
name: magnetics
description: Magnetic diagnostics for equilibrium identification and pl...
imas: magnetics
license_name: Creative Commons 4.0 BY-SA
license_url: https://creativecommons.org/licenses/by-sa/4.0/- b_field_pol_probe_cc_channel: 5
- time_mirnov: 363201
- b_field_pol_probe_cc_geometry_channel: 40
- b_field_pol_probe_ccbv_channel: 40
- time: 3633
- b_field_pol_probe_ccbv_geometry_channel: 40
- b_field_pol_probe_obr_channel: 19
- b_field_pol_probe_obr_geometry_channel: 19
- b_field_pol_probe_obv_channel: 19
- b_field_pol_probe_obv_geometry_channel: 19
- b_field_pol_probe_omv_geometry_channel: 21
- b_field_pol_probe_omv_channel: 3
- b_field_tor_probe_cc_channel: 3
- b_field_tor_probe_cc_geometry_channel: 36
- b_field_tor_probe_omaha_channel: 4
- time_omaha: 7264001
- b_field_tor_probe_saddle_field_channel: 12
- time_saddle: 36321
- b_field_tor_probe_saddle_l_geometry_channel: 12
- coordinate: 28
- b_field_tor_probe_saddle_m_geometry_channel: 12
- b_field_tor_probe_saddle_u_geometry_channel: 12
- b_field_tor_probe_saddle_voltage_channel: 12
- flux_loop_channel: 15
- flux_loop_geometry_channel: 44
- b_field_pol_probe_cc_channel(b_field_pol_probe_cc_channel)<U13'xmc/CC/MV/201' ... 'xmc/CC/MV/240'
array(['xmc/CC/MV/201', 'xmc/CC/MV/210', 'xmc/CC/MV/220', 'xmc/CC/MV/230', 'xmc/CC/MV/240'], dtype='<U13') - b_field_pol_probe_cc_geometry_channel(b_field_pol_probe_cc_geometry_channel)object'cc_mv_201' ... 'cc_mv_240'
array(['cc_mv_201', 'cc_mv_202*', 'cc_mv_203*', 'cc_mv_204*', 'cc_mv_205*', 'cc_mv_206*', 'cc_mv_207*', 'cc_mv_208*', 'cc_mv_209*', 'cc_mv_210*', 'cc_mv_211*', 'cc_mv_212*', 'cc_mv_213*', 'cc_mv_214*', 'cc_mv_215*', 'cc_mv_216*', 'cc_mv_217*', 'cc_mv_218*', 'cc_mv_219*', 'cc_mv_220*', 'cc_mv_221*', 'cc_mv_222*', 'cc_mv_223*', 'cc_mv_224', 'cc_mv_225', 'cc_mv_226', 'cc_mv_227', 'cc_mv_228', 'cc_mv_229', 'cc_mv_230', 'cc_mv_231', 'cc_mv_232', 'cc_mv_233', 'cc_mv_234', 'cc_mv_235', 'cc_mv_236', 'cc_mv_237', 'cc_mv_238', 'cc_mv_239', 'cc_mv_240'], dtype=object) - b_field_pol_probe_ccbv_channel(b_field_pol_probe_ccbv_channel)<U10'AMB_CCBV01' ... 'AMB_CCBV40'
array(['AMB_CCBV01', 'AMB_CCBV02', 'AMB_CCBV03', 'AMB_CCBV04', 'AMB_CCBV05', 'AMB_CCBV06', 'AMB_CCBV07', 'AMB_CCBV08', 'AMB_CCBV09', 'AMB_CCBV10', 'AMB_CCBV11', 'AMB_CCBV12', 'AMB_CCBV13', 'AMB_CCBV14', 'AMB_CCBV15', 'AMB_CCBV16', 'AMB_CCBV17', 'AMB_CCBV18', 'AMB_CCBV19', 'AMB_CCBV20', 'AMB_CCBV21', 'AMB_CCBV22', 'AMB_CCBV23', 'AMB_CCBV24', 'AMB_CCBV25', 'AMB_CCBV26', 'AMB_CCBV27', 'AMB_CCBV28', 'AMB_CCBV29', 'AMB_CCBV30', 'AMB_CCBV31', 'AMB_CCBV32', 'AMB_CCBV33', 'AMB_CCBV34', 'AMB_CCBV35', 'AMB_CCBV36', 'AMB_CCBV37', 'AMB_CCBV38', 'AMB_CCBV39', 'AMB_CCBV40'], dtype='<U10') - b_field_pol_probe_ccbv_geometry_channel(b_field_pol_probe_ccbv_geometry_channel)object'ccbv01' 'ccbv02' ... 'ccbv40'
array(['ccbv01', 'ccbv02', 'ccbv03', 'ccbv04', 'ccbv05', 'ccbv06', 'ccbv07', 'ccbv08', 'ccbv09', 'ccbv10', 'ccbv11', 'ccbv12', 'ccbv13', 'ccbv14', 'ccbv15', 'ccbv16', 'ccbv17', 'ccbv18', 'ccbv19', 'ccbv20', 'ccbv21', 'ccbv22', 'ccbv23', 'ccbv24', 'ccbv25', 'ccbv26', 'ccbv27', 'ccbv28', 'ccbv29', 'ccbv30', 'ccbv31', 'ccbv32', 'ccbv33', 'ccbv34', 'ccbv35', 'ccbv36', 'ccbv37', 'ccbv38', 'ccbv39', 'ccbv40'], dtype=object) - b_field_pol_probe_obr_channel(b_field_pol_probe_obr_channel)<U9'AMB_OBR01' ... 'AMB_OBR19'
array(['AMB_OBR01', 'AMB_OBR02', 'AMB_OBR03', 'AMB_OBR04', 'AMB_OBR05', 'AMB_OBR06', 'AMB_OBR07', 'AMB_OBR08', 'AMB_OBR09', 'AMB_OBR10', 'AMB_OBR11', 'AMB_OBR12', 'AMB_OBR13', 'AMB_OBR14', 'AMB_OBR15', 'AMB_OBR16', 'AMB_OBR17', 'AMB_OBR18', 'AMB_OBR19'], dtype='<U9') - b_field_pol_probe_obr_geometry_channel(b_field_pol_probe_obr_geometry_channel)object'obr01' 'obr02' ... 'obr18' 'obr19'
array(['obr01', 'obr02', 'obr03', 'obr04', 'obr05', 'obr06', 'obr07', 'obr08', 'obr09', 'obr10', 'obr11', 'obr12', 'obr13', 'obr14', 'obr15', 'obr16', 'obr17', 'obr18', 'obr19'], dtype=object) - b_field_pol_probe_obv_channel(b_field_pol_probe_obv_channel)<U9'AMB_OBV01' ... 'AMB_OBV19'
array(['AMB_OBV01', 'AMB_OBV02', 'AMB_OBV03', 'AMB_OBV04', 'AMB_OBV05', 'AMB_OBV06', 'AMB_OBV07', 'AMB_OBV08', 'AMB_OBV09', 'AMB_OBV10', 'AMB_OBV11', 'AMB_OBV12', 'AMB_OBV13', 'AMB_OBV14', 'AMB_OBV15', 'AMB_OBV16', 'AMB_OBV17', 'AMB_OBV18', 'AMB_OBV19'], dtype='<U9') - b_field_pol_probe_obv_geometry_channel(b_field_pol_probe_obv_geometry_channel)object'obv01' 'obv02' ... 'obv18' 'obv19'
array(['obv01', 'obv02', 'obv03', 'obv04', 'obv05', 'obv06', 'obv07', 'obv08', 'obv09', 'obv10', 'obv11', 'obv12', 'obv13', 'obv14', 'obv15', 'obv16', 'obv17', 'obv18', 'obv19'], dtype=object) - b_field_pol_probe_omv_channel(b_field_pol_probe_omv_channel)<U11'xmc/OMV/110' ... 'xmc/OMV/310'
array(['xmc/OMV/110', 'xmc/OMV/210', 'xmc/OMV/310'], dtype='<U11')
- b_field_pol_probe_omv_geometry_channel(b_field_pol_probe_omv_geometry_channel)object'omv_201' 'omv_202' ... 'omv_310'
array(['omv_201', 'omv_202', 'omv_203', 'omv_204', 'omv_205', 'omv_206', 'omv_207', 'omv_208', 'omv_209', 'omv_210', 'omv_211', 'omv_212', 'omv_213', 'omv_214', 'omv_215', 'omv_216', 'omv_217', 'omv_218', 'omv_219', 'omv_110', 'omv_310'], dtype=object) - b_field_tor_probe_cc_channel(b_field_tor_probe_cc_channel)<U13'xmc/CC/MT/201' ... 'xmc/CC/MT/212'
array(['xmc/CC/MT/201', 'xmc/CC/MT/206', 'xmc/CC/MT/212'], dtype='<U13')
- b_field_tor_probe_cc_geometry_channel(b_field_tor_probe_cc_geometry_channel)object'cc_mt_101' ... 'cc_mt_312'
array(['cc_mt_101', 'cc_mt_102', 'cc_mt_103', 'cc_mt_104', 'cc_mt_105', 'cc_mt_106', 'cc_mt_107', 'cc_mt_108', 'cc_mt_109', 'cc_mt_110', 'cc_mt_111', 'cc_mt_112', 'cc_mt_201', 'cc_mt_202', 'cc_mt_203', 'cc_mt_204', 'cc_mt_205', 'cc_mt_206', 'cc_mt_207', 'cc_mt_208', 'cc_mt_209', 'cc_mt_210', 'cc_mt_211', 'cc_mt_212', 'cc_mt_301', 'cc_mt_302', 'cc_mt_303', 'cc_mt_304', 'cc_mt_305', 'cc_mt_306', 'cc_mt_307', 'cc_mt_308', 'cc_mt_309', 'cc_mt_310', 'cc_mt_311', 'cc_mt_312'], dtype=object) - b_field_tor_probe_omaha_channel(b_field_tor_probe_omaha_channel)<U14'/xmo/OMAHA/1LZ' ... '/xmo/OMAHA...
array(['/xmo/OMAHA/1LZ', '/xmo/OMAHA/3LZ', '/xmo/OMAHA/5LZ', '/xmo/OMAHA/6LZ'], dtype='<U14') - b_field_tor_probe_saddle_field_channel(b_field_tor_probe_saddle_field_channel)<U11'ASM_SAD/M01' ... 'ASM_SAD/M12'
array(['ASM_SAD/M01', 'ASM_SAD/M02', 'ASM_SAD/M03', 'ASM_SAD/M04', 'ASM_SAD/M05', 'ASM_SAD/M06', 'ASM_SAD/M07', 'ASM_SAD/M08', 'ASM_SAD/M09', 'ASM_SAD/M10', 'ASM_SAD/M11', 'ASM_SAD/M12'], dtype='<U11') - b_field_tor_probe_saddle_l_geometry_channel(b_field_tor_probe_saddle_l_geometry_channel)object'sad_out_l01' ... 'sad_out_l12'
array(['sad_out_l01', 'sad_out_l02', 'sad_out_l03', 'sad_out_l04', 'sad_out_l05', 'sad_out_l06', 'sad_out_l07', 'sad_out_l08', 'sad_out_l09', 'sad_out_l10', 'sad_out_l11', 'sad_out_l12'], dtype=object) - b_field_tor_probe_saddle_m_geometry_channel(b_field_tor_probe_saddle_m_geometry_channel)object'sad_out_m01' ... 'sad_out_m12'
array(['sad_out_m01', 'sad_out_m02', 'sad_out_m03', 'sad_out_m04', 'sad_out_m05', 'sad_out_m06', 'sad_out_m07', 'sad_out_m08', 'sad_out_m09', 'sad_out_m10', 'sad_out_m11', 'sad_out_m12'], dtype=object) - b_field_tor_probe_saddle_u_geometry_channel(b_field_tor_probe_saddle_u_geometry_channel)object'sad_out_u01' ... 'sad_out_u12'
array(['sad_out_u01', 'sad_out_u02', 'sad_out_u03', 'sad_out_u04', 'sad_out_u05', 'sad_out_u06', 'sad_out_u07', 'sad_out_u08', 'sad_out_u09', 'sad_out_u10', 'sad_out_u11', 'sad_out_u12'], dtype=object) - b_field_tor_probe_saddle_voltage_channel(b_field_tor_probe_saddle_voltage_channel)<U15'XMB/SAD/OUT/M01' ... 'XMB/SAD/O...
array(['XMB/SAD/OUT/M01', 'XMB/SAD/OUT/M02', 'XMB/SAD/OUT/M03', 'XMB/SAD/OUT/M04', 'XMB/SAD/OUT/M05', 'XMB/SAD/OUT/M06', 'XMB/SAD/OUT/M07', 'XMB/SAD/OUT/M08', 'XMB/SAD/OUT/M09', 'XMB/SAD/OUT/M10', 'XMB/SAD/OUT/M11', 'XMB/SAD/OUT/M12'], dtype='<U15') - coordinate(coordinate)int640 1 2 3 4 5 6 ... 22 23 24 25 26 27
array([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27]) - flux_loop_channel(flux_loop_channel)<U12'AMB_FL/CC03' ... 'AMB_FL/P6L/1'
array(['AMB_FL/CC03', 'AMB_FL/CC04', 'AMB_FL/CC05', 'AMB_FL/CC07', 'AMB_FL/CC09', 'AMB_FL/P3L/4', 'AMB_FL/P3U/1', 'AMB_FL/P3U/4', 'AMB_FL/P4L/1', 'AMB_FL/P4L/4', 'AMB_FL/P4U/4', 'AMB_FL/P5L/1', 'AMB_FL/P5L/4', 'AMB_FL/P5U/1', 'AMB_FL/P6L/1'], dtype='<U12') - flux_loop_geometry_channel(flux_loop_geometry_channel)object'FL_P2U_1' ... 'FL_CC010'
array(['FL_P2U_1', 'FL_P2U_2', 'FL_P2U_3', 'FL_P2U_4', 'FL_P2L_1', 'FL_P2L_2', 'FL_P2L_3', 'FL_P2L_4', 'FL_P3U_1', 'FL_P3U_2', 'FL_P3U_3', 'FL_P3U_4', 'FL_P3L_1', 'FL_P3L_2', 'FL_P3L_3', 'FL_P3L_4', 'FL_P4U_1', 'FL_P4U_2', 'FL_P4U_3', 'FL_P4U_4', 'FL_P4L_1', 'FL_P4L_2', 'FL_P4L_3', 'FL_P4L_4', 'FL_P5U_1', 'FL_P5U_2', 'FL_P5U_3', 'FL_P5U_4', 'FL_P5L_1', 'FL_P5L_2', 'FL_P5L_3', 'FL_P5L_4', 'FL_P6L_1', 'FL_P6L_2', 'FL_CC01', 'FL_CC02', 'FL_CC03', 'FL_CC04', 'FL_CC05', 'FL_CC06', 'FL_CC07', 'FL_CC08', 'FL_CC09', 'FL_CC010'], dtype=object) - time(time)float64-0.0612 -0.061 ... 0.665 0.6652
- units :
- s
array([-0.0612, -0.061 , -0.0608, ..., 0.6648, 0.665 , 0.6652], shape=(3633,)) - time_mirnov(time_mirnov)float64-0.0612 -0.0612 ... 0.6652 0.6652
- units :
- s
array([-0.0612 , -0.061198, -0.061196, ..., 0.665196, 0.665198, 0.6652 ], shape=(363201,)) - time_omaha(time_omaha)float64-0.0612 -0.0612 ... 0.6652 0.6652
- units :
- s
array([-0.0612, -0.0612, -0.0612, ..., 0.6652, 0.6652, 0.6652], shape=(7264001,)) - time_saddle(time_saddle)float64-0.0612 -0.06118 ... 0.6652 0.6652
- units :
- s
array([-0.0612 , -0.06118, -0.06116, ..., 0.66516, 0.66518, 0.6652 ], shape=(36321,))
- b_field_pol_probe_cc_field(b_field_pol_probe_cc_channel, time_mirnov)float64...
- data_block_size :
- 5314444
- description :
- Centre column poloidal mirnov array
- label :
- Tesla/sec
- name :
- b_field_pol_probe_cc_field
- uda_name :
- xmc/CC/MV/201
- imas :
- magnetics.b_field_pol_probe[:].field.data
- units :
- T
[1816005 values with dtype=float64]
- b_field_pol_probe_cc_phi(b_field_pol_probe_cc_geometry_channel)float64...
- device :
- MAST
- class_ :
- magnetics
- system :
- mirnov
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the mirnov coils for MAST
- comment :
- Data sourced from configuration PDF
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_mirnovs.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-08
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/XMC_OMV - Outboard vertical (Bv_Br) Mirnov Arrays.pdf, https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/CCMV - Centre Column Vertical Array of Bv Mirnovs.pdf, https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/CCMT - CC toroidal Arrays of Bv Mirnovs.pdf
- imas :
- description :
- Toroidal angle of the centre column poloidal mirnov array
[40 values with dtype=float64]
- b_field_pol_probe_cc_r(b_field_pol_probe_cc_geometry_channel)float64...
- device :
- MAST
- class_ :
- magnetics
- system :
- mirnov
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the mirnov coils for MAST
- comment :
- Data sourced from configuration PDF
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_mirnovs.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-08
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/XMC_OMV - Outboard vertical (Bv_Br) Mirnov Arrays.pdf, https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/CCMV - Centre Column Vertical Array of Bv Mirnovs.pdf, https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/CCMT - CC toroidal Arrays of Bv Mirnovs.pdf
- imas :
- description :
- Major radius of the centre column poloidal mirnov array
[40 values with dtype=float64]
- b_field_pol_probe_cc_z(b_field_pol_probe_cc_geometry_channel)float64...
- device :
- MAST
- class_ :
- magnetics
- system :
- mirnov
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the mirnov coils for MAST
- comment :
- Data sourced from configuration PDF
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_mirnovs.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-08
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/XMC_OMV - Outboard vertical (Bv_Br) Mirnov Arrays.pdf, https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/CCMV - Centre Column Vertical Array of Bv Mirnovs.pdf, https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/CCMT - CC toroidal Arrays of Bv Mirnovs.pdf
- imas :
- description :
- Vertical position of the centre column poloidal mirnov array
[40 values with dtype=float64]
- b_field_pol_probe_ccbv_field(b_field_pol_probe_ccbv_channel, time)float64...
- data_block_size :
- 390428
- description :
- Centre column Bv array
- label :
- CCBV01
- name :
- b_field_pol_probe_ccbv_field
- uda_name :
- AMB_CCBV01
- imas :
- magnetics.b_field_pol_probe[:].field.data
- units :
- T
[145320 values with dtype=float64]
- b_field_pol_probe_ccbv_length(b_field_pol_probe_ccbv_geometry_channel)float64...
- device :
- MAST
- class_ :
- magnetics
- system :
- pickup
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the pickup coils for MAST
- comment :
- Data sourced from configuration PDF
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_pickup.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-08-15
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/XMA_OBV_OBR Outer Bv_Br Discrete Arrays.pdf, https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/Centre Column Vertical Bv Arrays - XMA_CCBV.pdf
- imas :
- description :
- Length of the centre column Bv array
[40 values with dtype=float64]
- b_field_pol_probe_ccbv_phi_1(b_field_pol_probe_ccbv_geometry_channel)float64...
- device :
- MAST
- class_ :
- magnetics
- system :
- pickup
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the pickup coils for MAST
- comment :
- Data sourced from configuration PDF
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_pickup.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-08-15
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/XMA_OBV_OBR Outer Bv_Br Discrete Arrays.pdf, https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/Centre Column Vertical Bv Arrays - XMA_CCBV.pdf
- imas :
- description :
- Phi 1 of the centre column Bv array
[40 values with dtype=float64]
- b_field_pol_probe_ccbv_phi_2(b_field_pol_probe_ccbv_geometry_channel)float64...
- device :
- MAST
- class_ :
- magnetics
- system :
- pickup
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the pickup coils for MAST
- comment :
- Data sourced from configuration PDF
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_pickup.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-08-15
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/XMA_OBV_OBR Outer Bv_Br Discrete Arrays.pdf, https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/Centre Column Vertical Bv Arrays - XMA_CCBV.pdf
- imas :
- description :
- Phi 2 of the centre column Bv array
[40 values with dtype=float64]
- b_field_pol_probe_ccbv_r(b_field_pol_probe_ccbv_geometry_channel)float64...
- device :
- MAST
- class_ :
- magnetics
- system :
- pickup
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the pickup coils for MAST
- comment :
- Data sourced from configuration PDF
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_pickup.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-08-15
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/XMA_OBV_OBR Outer Bv_Br Discrete Arrays.pdf, https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/Centre Column Vertical Bv Arrays - XMA_CCBV.pdf
- imas :
- description :
- Major radius of the centre column Bv array
[40 values with dtype=float64]
- b_field_pol_probe_ccbv_z(b_field_pol_probe_ccbv_geometry_channel)float64...
- device :
- MAST
- class_ :
- magnetics
- system :
- pickup
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the pickup coils for MAST
- comment :
- Data sourced from configuration PDF
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_pickup.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-08-15
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/XMA_OBV_OBR Outer Bv_Br Discrete Arrays.pdf, https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/Centre Column Vertical Bv Arrays - XMA_CCBV.pdf
- imas :
- description :
- Vertical position of the centre column Bv array
[40 values with dtype=float64]
- b_field_pol_probe_obr_field(b_field_pol_probe_obr_channel, time)float64...
- data_block_size :
- 474428
- description :
- Outer Br array
- label :
- OBR01
- name :
- b_field_pol_probe_obr_field
- uda_name :
- AMB_OBR01
- imas :
- magnetics.b_field_pol_probe[:].field.data
- units :
- T
[69027 values with dtype=float64]
- b_field_pol_probe_obr_length(b_field_pol_probe_obr_geometry_channel)float64...
- device :
- MAST
- class_ :
- magnetics
- system :
- pickup
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the pickup coils for MAST
- comment :
- Data sourced from configuration PDF
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_pickup.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-08-15
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/XMA_OBV_OBR Outer Bv_Br Discrete Arrays.pdf, https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/Centre Column Vertical Bv Arrays - XMA_CCBV.pdf
- imas :
- description :
- Length of the outer vessel Br array
[19 values with dtype=float64]
- b_field_pol_probe_obr_phi_1(b_field_pol_probe_obr_geometry_channel)float64...
- device :
- MAST
- class_ :
- magnetics
- system :
- pickup
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the pickup coils for MAST
- comment :
- Data sourced from configuration PDF
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_pickup.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-08-15
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/XMA_OBV_OBR Outer Bv_Br Discrete Arrays.pdf, https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/Centre Column Vertical Bv Arrays - XMA_CCBV.pdf
- imas :
- description :
- Phi 1 of the outer vessel Br array
[19 values with dtype=float64]
- b_field_pol_probe_obr_phi_2(b_field_pol_probe_obr_geometry_channel)float64...
- device :
- MAST
- class_ :
- magnetics
- system :
- pickup
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the pickup coils for MAST
- comment :
- Data sourced from configuration PDF
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_pickup.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-08-15
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/XMA_OBV_OBR Outer Bv_Br Discrete Arrays.pdf, https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/Centre Column Vertical Bv Arrays - XMA_CCBV.pdf
- imas :
- description :
- Phi 2 of the outer vessel Br array
[19 values with dtype=float64]
- b_field_pol_probe_obr_r(b_field_pol_probe_obr_geometry_channel)float64...
- device :
- MAST
- class_ :
- magnetics
- system :
- pickup
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the pickup coils for MAST
- comment :
- Data sourced from configuration PDF
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_pickup.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-08-15
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/XMA_OBV_OBR Outer Bv_Br Discrete Arrays.pdf, https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/Centre Column Vertical Bv Arrays - XMA_CCBV.pdf
- imas :
- description :
- Major radius of the outer vessel Br array
[19 values with dtype=float64]
- b_field_pol_probe_obr_z(b_field_pol_probe_obr_geometry_channel)float64...
- device :
- MAST
- class_ :
- magnetics
- system :
- pickup
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the pickup coils for MAST
- comment :
- Data sourced from configuration PDF
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_pickup.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-08-15
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/XMA_OBV_OBR Outer Bv_Br Discrete Arrays.pdf, https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/Centre Column Vertical Bv Arrays - XMA_CCBV.pdf
- imas :
- description :
- Vertical position of the outer vessel Br array
[19 values with dtype=float64]
- b_field_pol_probe_obv_field(b_field_pol_probe_obv_channel, time)float64...
- imas :
- magnetics.b_field_pol_probe[:].field.data
- description :
- Outer Bv array
- name :
- b_field_pol_probe_obv_field
- units :
- T
[69027 values with dtype=float64]
- b_field_pol_probe_obv_length(b_field_pol_probe_obv_geometry_channel)float64...
- device :
- MAST
- class_ :
- magnetics
- system :
- pickup
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the pickup coils for MAST
- comment :
- Data sourced from configuration PDF
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_pickup.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-08-15
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/XMA_OBV_OBR Outer Bv_Br Discrete Arrays.pdf, https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/Centre Column Vertical Bv Arrays - XMA_CCBV.pdf
- imas :
- description :
- Length of the outer vessel Bv array
[19 values with dtype=float64]
- b_field_pol_probe_obv_phi_1(b_field_pol_probe_obv_geometry_channel)float64...
- device :
- MAST
- class_ :
- magnetics
- system :
- pickup
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the pickup coils for MAST
- comment :
- Data sourced from configuration PDF
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_pickup.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-08-15
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/XMA_OBV_OBR Outer Bv_Br Discrete Arrays.pdf, https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/Centre Column Vertical Bv Arrays - XMA_CCBV.pdf
- imas :
- description :
- Phi 1 of the outer vessel Bv array
[19 values with dtype=float64]
- b_field_pol_probe_obv_phi_2(b_field_pol_probe_obv_geometry_channel)float64...
- device :
- MAST
- class_ :
- magnetics
- system :
- pickup
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the pickup coils for MAST
- comment :
- Data sourced from configuration PDF
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_pickup.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-08-15
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/XMA_OBV_OBR Outer Bv_Br Discrete Arrays.pdf, https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/Centre Column Vertical Bv Arrays - XMA_CCBV.pdf
- imas :
- description :
- Phi 2 of the outer vessel Bv array
[19 values with dtype=float64]
- b_field_pol_probe_obv_r(b_field_pol_probe_obv_geometry_channel)float64...
- device :
- MAST
- class_ :
- magnetics
- system :
- pickup
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the pickup coils for MAST
- comment :
- Data sourced from configuration PDF
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_pickup.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-08-15
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/XMA_OBV_OBR Outer Bv_Br Discrete Arrays.pdf, https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/Centre Column Vertical Bv Arrays - XMA_CCBV.pdf
- imas :
- description :
- Major radius of the outer vessel Bv array
[19 values with dtype=float64]
- b_field_pol_probe_obv_z(b_field_pol_probe_obv_geometry_channel)float64...
- device :
- MAST
- class_ :
- magnetics
- system :
- pickup
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the pickup coils for MAST
- comment :
- Data sourced from configuration PDF
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_pickup.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-08-15
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/XMA_OBV_OBR Outer Bv_Br Discrete Arrays.pdf, https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/Centre Column Vertical Bv Arrays - XMA_CCBV.pdf
- imas :
- description :
- Vertical position of the outer vessel Bv array
[19 values with dtype=float64]
- b_field_pol_probe_omv_phi(b_field_pol_probe_omv_geometry_channel)float64...
- device :
- MAST
- class_ :
- magnetics
- system :
- mirnov
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the mirnov coils for MAST
- comment :
- Data sourced from configuration PDF
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_mirnovs.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-08
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/XMC_OMV - Outboard vertical (Bv_Br) Mirnov Arrays.pdf, https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/CCMV - Centre Column Vertical Array of Bv Mirnovs.pdf, https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/CCMT - CC toroidal Arrays of Bv Mirnovs.pdf
- imas :
- description :
- Toroidal angle of the Outboard vertical mirnov array
[21 values with dtype=float64]
- b_field_pol_probe_omv_r(b_field_pol_probe_omv_geometry_channel)float64...
- device :
- MAST
- class_ :
- magnetics
- system :
- mirnov
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the mirnov coils for MAST
- comment :
- Data sourced from configuration PDF
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_mirnovs.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-08
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/XMC_OMV - Outboard vertical (Bv_Br) Mirnov Arrays.pdf, https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/CCMV - Centre Column Vertical Array of Bv Mirnovs.pdf, https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/CCMT - CC toroidal Arrays of Bv Mirnovs.pdf
- imas :
- description :
- Major radius of the Outboard vertical mirnov array
[21 values with dtype=float64]
- b_field_pol_probe_omv_voltage(b_field_pol_probe_omv_channel, time_mirnov)float64...
- data_block_size :
- 5314444
- description :
- Outboard vertical mirnov array
- label :
- Volt
- name :
- b_field_pol_probe_omv_voltage
- uda_name :
- xmc/OMV/110
- imas :
- magnetics.b_field_tor_probe[:].voltage.data
- units :
- V
[1089603 values with dtype=float64]
- b_field_pol_probe_omv_z(b_field_pol_probe_omv_geometry_channel)float64...
- device :
- MAST
- class_ :
- magnetics
- system :
- mirnov
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the mirnov coils for MAST
- comment :
- Data sourced from configuration PDF
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_mirnovs.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-08
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/XMC_OMV - Outboard vertical (Bv_Br) Mirnov Arrays.pdf, https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/CCMV - Centre Column Vertical Array of Bv Mirnovs.pdf, https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/CCMT - CC toroidal Arrays of Bv Mirnovs.pdf
- imas :
- description :
- Vertical position of the Outboard vertical mirnov array
[21 values with dtype=float64]
- b_field_tor_probe_cc_field(b_field_tor_probe_cc_channel, time_mirnov)float64...
- data_block_size :
- 5314444
- description :
- Centre column toroidal mirnov array
- label :
- Tesla/sec
- name :
- b_field_tor_probe_cc_field
- uda_name :
- xmc/CC/MT/201
- imas :
- magnetics.b_field_tor_probe[:].field.data
- units :
- T
[1089603 values with dtype=float64]
- b_field_tor_probe_cc_phi(b_field_tor_probe_cc_geometry_channel)float64...
- device :
- MAST
- class_ :
- magnetics
- system :
- mirnov
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the mirnov coils for MAST
- comment :
- Data sourced from configuration PDF
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_mirnovs.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-08
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/XMC_OMV - Outboard vertical (Bv_Br) Mirnov Arrays.pdf, https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/CCMV - Centre Column Vertical Array of Bv Mirnovs.pdf, https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/CCMT - CC toroidal Arrays of Bv Mirnovs.pdf
- imas :
- description :
- Toroidal angle of the centre column toroidal mirnov array
[36 values with dtype=float64]
- b_field_tor_probe_cc_r(b_field_tor_probe_cc_geometry_channel)float64...
- device :
- MAST
- class_ :
- magnetics
- system :
- mirnov
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the mirnov coils for MAST
- comment :
- Data sourced from configuration PDF
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_mirnovs.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-08
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/XMC_OMV - Outboard vertical (Bv_Br) Mirnov Arrays.pdf, https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/CCMV - Centre Column Vertical Array of Bv Mirnovs.pdf, https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/CCMT - CC toroidal Arrays of Bv Mirnovs.pdf
- imas :
- description :
- Major radius of the centre column toroidal mirnov array
[36 values with dtype=float64]
- b_field_tor_probe_cc_z(b_field_tor_probe_cc_geometry_channel)float64...
- device :
- MAST
- class_ :
- magnetics
- system :
- mirnov
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the mirnov coils for MAST
- comment :
- Data sourced from configuration PDF
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_mirnovs.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-08
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/XMC_OMV - Outboard vertical (Bv_Br) Mirnov Arrays.pdf, https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/CCMV - Centre Column Vertical Array of Bv Mirnovs.pdf, https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/CCMT - CC toroidal Arrays of Bv Mirnovs.pdf
- imas :
- description :
- Vertical position of the centre column toroidal mirnov array
[36 values with dtype=float64]
- b_field_tor_probe_omaha_voltage(b_field_tor_probe_omaha_channel, time_omaha)float64...
- data_block_size :
- 56114444
- description :
- High frequency OMAHA toroidal mirnov array
- label :
- arb
- name :
- b_field_tor_probe_omaha_voltage
- uda_name :
- /xmo/OMAHA/1LZ
- imas :
- magnetics.b_field_tor_probe[:].voltage.data
- units :
- V
[29056004 values with dtype=float64]
- b_field_tor_probe_saddle_field(b_field_tor_probe_saddle_field_channel, time_saddle)float64...
- data_block_size :
- 150620
- description :
- label :
- mT
- name :
- b_field_tor_probe_saddle_field
- uda_name :
- ASM_SAD/M01
- imas :
- magnetics.b_field_tor_probe[:].field.data
- units :
- T
[435852 values with dtype=float64]
- b_field_tor_probe_saddle_l_phi(b_field_tor_probe_saddle_l_geometry_channel, coordinate)float64...
- device :
- MAST
- class_ :
- magnetics
- system :
- saddle coils
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the saddle coils for MAST
- comment :
- Data sourced from configuration PDF
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_saddle.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-01
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/XMB_SAD Outer Saddle Coils.pdf
- imas :
- description :
- Toroidal points of the lower saddle coils
[336 values with dtype=float64]
- b_field_tor_probe_saddle_l_r(b_field_tor_probe_saddle_l_geometry_channel, coordinate)float64...
- device :
- MAST
- class_ :
- magnetics
- system :
- saddle coils
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the saddle coils for MAST
- comment :
- Data sourced from configuration PDF
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_saddle.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-01
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/XMB_SAD Outer Saddle Coils.pdf
- imas :
- description :
- Radial points of the lower saddle coils
[336 values with dtype=float64]
- b_field_tor_probe_saddle_l_z(b_field_tor_probe_saddle_l_geometry_channel, coordinate)float64...
- device :
- MAST
- class_ :
- magnetics
- system :
- saddle coils
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the saddle coils for MAST
- comment :
- Data sourced from configuration PDF
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_saddle.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-01
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/XMB_SAD Outer Saddle Coils.pdf
- imas :
- description :
- Vertical positions of the lower saddle coils
[336 values with dtype=float64]
- b_field_tor_probe_saddle_m_phi(b_field_tor_probe_saddle_m_geometry_channel, coordinate)float64...
- device :
- MAST
- class_ :
- magnetics
- system :
- saddle coils
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the saddle coils for MAST
- comment :
- Data sourced from configuration PDF
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_saddle.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-01
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/XMB_SAD Outer Saddle Coils.pdf
- imas :
- description :
- Toroidal points of the middle saddle coils
[336 values with dtype=float64]
- b_field_tor_probe_saddle_m_r(b_field_tor_probe_saddle_m_geometry_channel, coordinate)float64...
- device :
- MAST
- class_ :
- magnetics
- system :
- saddle coils
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the saddle coils for MAST
- comment :
- Data sourced from configuration PDF
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_saddle.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-01
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/XMB_SAD Outer Saddle Coils.pdf
- imas :
- description :
- Radial points of the middle saddle coils
[336 values with dtype=float64]
- b_field_tor_probe_saddle_m_z(b_field_tor_probe_saddle_m_geometry_channel, coordinate)float64...
- device :
- MAST
- class_ :
- magnetics
- system :
- saddle coils
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the saddle coils for MAST
- comment :
- Data sourced from configuration PDF
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_saddle.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-01
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/XMB_SAD Outer Saddle Coils.pdf
- imas :
- description :
- Vertical positions of the middle saddle coils
[336 values with dtype=float64]
- b_field_tor_probe_saddle_u_phi(b_field_tor_probe_saddle_u_geometry_channel, coordinate)float64...
- device :
- MAST
- class_ :
- magnetics
- system :
- saddle coils
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the saddle coils for MAST
- comment :
- Data sourced from configuration PDF
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_saddle.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-01
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/XMB_SAD Outer Saddle Coils.pdf
- imas :
- description :
- Toroidal points of the upper saddle coils
[336 values with dtype=float64]
- b_field_tor_probe_saddle_u_r(b_field_tor_probe_saddle_u_geometry_channel, coordinate)float64...
- device :
- MAST
- class_ :
- magnetics
- system :
- saddle coils
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the saddle coils for MAST
- comment :
- Data sourced from configuration PDF
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_saddle.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-01
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/XMB_SAD Outer Saddle Coils.pdf
- imas :
- description :
- Radial points of the upper saddle coils
[336 values with dtype=float64]
- b_field_tor_probe_saddle_u_z(b_field_tor_probe_saddle_u_geometry_channel, coordinate)float64...
- device :
- MAST
- class_ :
- magnetics
- system :
- saddle coils
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the saddle coils for MAST
- comment :
- Data sourced from configuration PDF
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_saddle.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-01
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/XMB_SAD Outer Saddle Coils.pdf
- imas :
- description :
- Vertical positions of the upper saddle coils
[336 values with dtype=float64]
- b_field_tor_probe_saddle_voltage(b_field_tor_probe_saddle_voltage_channel, time_saddle)float64...
- data_block_size :
- 774444
- description :
- label :
- V
- name :
- b_field_tor_probe_saddle_voltage
- uda_name :
- XMB/SAD/OUT/M01
- imas :
- magnetics.b_field_tor_probe[:].voltage.data
- units :
- V
[435852 values with dtype=float64]
- flux_loop_flux(flux_loop_channel, time)float64...
- data_block_size :
- 474428
- description :
- label :
- FL/CC03
- name :
- flux_loop_flux
- uda_name :
- AMB_FL/CC03
- imas :
- magnetics.flux_loop[:].flux.data
- units :
- Wb
[54495 values with dtype=float64]
- flux_loop_r(flux_loop_geometry_channel)float64...
- device :
- MAST
- class_ :
- magnetics
- system :
- fluxloops
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the fluxloops for MAST
- comment :
- Data sourced from EFIT file
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_fluxloops.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-08
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/detectors.dat_M4
- imas :
- description :
- Major radius of the flux loops
[44 values with dtype=float64]
- flux_loop_z(flux_loop_geometry_channel)float64...
- device :
- MAST
- class_ :
- magnetics
- system :
- fluxloops
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the fluxloops for MAST
- comment :
- Data sourced from EFIT file
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_fluxloops.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-08
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/detectors.dat_M4
- imas :
- description :
- Vertical position of the flux loops
[44 values with dtype=float64]
- ip(time)float64-6.039e+03 -6.039e+03 ... 427.5
- data_block_size :
- 474428
- description :
- label :
- Plasma Current
- name :
- ip
- uda_name :
- AMC_PLASMA CURRENT
- imas :
- magnetics.ip[:].data
- units :
- A
array([-6039.37793 , -6039.37793 , -6516.910645, ..., 617.256775, 427.544525, 427.544525], shape=(3633,))
- b_field_pol_probe_cc_channelPandasIndex
PandasIndex(Index(['xmc/CC/MV/201', 'xmc/CC/MV/210', 'xmc/CC/MV/220', 'xmc/CC/MV/230', 'xmc/CC/MV/240'], dtype='object', name='b_field_pol_probe_cc_channel')) - b_field_pol_probe_cc_geometry_channelPandasIndex
PandasIndex(Index(['cc_mv_201', 'cc_mv_202*', 'cc_mv_203*', 'cc_mv_204*', 'cc_mv_205*', 'cc_mv_206*', 'cc_mv_207*', 'cc_mv_208*', 'cc_mv_209*', 'cc_mv_210*', 'cc_mv_211*', 'cc_mv_212*', 'cc_mv_213*', 'cc_mv_214*', 'cc_mv_215*', 'cc_mv_216*', 'cc_mv_217*', 'cc_mv_218*', 'cc_mv_219*', 'cc_mv_220*', 'cc_mv_221*', 'cc_mv_222*', 'cc_mv_223*', 'cc_mv_224', 'cc_mv_225', 'cc_mv_226', 'cc_mv_227', 'cc_mv_228', 'cc_mv_229', 'cc_mv_230', 'cc_mv_231', 'cc_mv_232', 'cc_mv_233', 'cc_mv_234', 'cc_mv_235', 'cc_mv_236', 'cc_mv_237', 'cc_mv_238', 'cc_mv_239', 'cc_mv_240'], dtype='object', name='b_field_pol_probe_cc_geometry_channel')) - b_field_pol_probe_ccbv_channelPandasIndex
PandasIndex(Index(['AMB_CCBV01', 'AMB_CCBV02', 'AMB_CCBV03', 'AMB_CCBV04', 'AMB_CCBV05', 'AMB_CCBV06', 'AMB_CCBV07', 'AMB_CCBV08', 'AMB_CCBV09', 'AMB_CCBV10', 'AMB_CCBV11', 'AMB_CCBV12', 'AMB_CCBV13', 'AMB_CCBV14', 'AMB_CCBV15', 'AMB_CCBV16', 'AMB_CCBV17', 'AMB_CCBV18', 'AMB_CCBV19', 'AMB_CCBV20', 'AMB_CCBV21', 'AMB_CCBV22', 'AMB_CCBV23', 'AMB_CCBV24', 'AMB_CCBV25', 'AMB_CCBV26', 'AMB_CCBV27', 'AMB_CCBV28', 'AMB_CCBV29', 'AMB_CCBV30', 'AMB_CCBV31', 'AMB_CCBV32', 'AMB_CCBV33', 'AMB_CCBV34', 'AMB_CCBV35', 'AMB_CCBV36', 'AMB_CCBV37', 'AMB_CCBV38', 'AMB_CCBV39', 'AMB_CCBV40'], dtype='object', name='b_field_pol_probe_ccbv_channel')) - b_field_pol_probe_ccbv_geometry_channelPandasIndex
PandasIndex(Index(['ccbv01', 'ccbv02', 'ccbv03', 'ccbv04', 'ccbv05', 'ccbv06', 'ccbv07', 'ccbv08', 'ccbv09', 'ccbv10', 'ccbv11', 'ccbv12', 'ccbv13', 'ccbv14', 'ccbv15', 'ccbv16', 'ccbv17', 'ccbv18', 'ccbv19', 'ccbv20', 'ccbv21', 'ccbv22', 'ccbv23', 'ccbv24', 'ccbv25', 'ccbv26', 'ccbv27', 'ccbv28', 'ccbv29', 'ccbv30', 'ccbv31', 'ccbv32', 'ccbv33', 'ccbv34', 'ccbv35', 'ccbv36', 'ccbv37', 'ccbv38', 'ccbv39', 'ccbv40'], dtype='object', name='b_field_pol_probe_ccbv_geometry_channel')) - b_field_pol_probe_obr_channelPandasIndex
PandasIndex(Index(['AMB_OBR01', 'AMB_OBR02', 'AMB_OBR03', 'AMB_OBR04', 'AMB_OBR05', 'AMB_OBR06', 'AMB_OBR07', 'AMB_OBR08', 'AMB_OBR09', 'AMB_OBR10', 'AMB_OBR11', 'AMB_OBR12', 'AMB_OBR13', 'AMB_OBR14', 'AMB_OBR15', 'AMB_OBR16', 'AMB_OBR17', 'AMB_OBR18', 'AMB_OBR19'], dtype='object', name='b_field_pol_probe_obr_channel')) - b_field_pol_probe_obr_geometry_channelPandasIndex
PandasIndex(Index(['obr01', 'obr02', 'obr03', 'obr04', 'obr05', 'obr06', 'obr07', 'obr08', 'obr09', 'obr10', 'obr11', 'obr12', 'obr13', 'obr14', 'obr15', 'obr16', 'obr17', 'obr18', 'obr19'], dtype='object', name='b_field_pol_probe_obr_geometry_channel')) - b_field_pol_probe_obv_channelPandasIndex
PandasIndex(Index(['AMB_OBV01', 'AMB_OBV02', 'AMB_OBV03', 'AMB_OBV04', 'AMB_OBV05', 'AMB_OBV06', 'AMB_OBV07', 'AMB_OBV08', 'AMB_OBV09', 'AMB_OBV10', 'AMB_OBV11', 'AMB_OBV12', 'AMB_OBV13', 'AMB_OBV14', 'AMB_OBV15', 'AMB_OBV16', 'AMB_OBV17', 'AMB_OBV18', 'AMB_OBV19'], dtype='object', name='b_field_pol_probe_obv_channel')) - b_field_pol_probe_obv_geometry_channelPandasIndex
PandasIndex(Index(['obv01', 'obv02', 'obv03', 'obv04', 'obv05', 'obv06', 'obv07', 'obv08', 'obv09', 'obv10', 'obv11', 'obv12', 'obv13', 'obv14', 'obv15', 'obv16', 'obv17', 'obv18', 'obv19'], dtype='object', name='b_field_pol_probe_obv_geometry_channel')) - b_field_pol_probe_omv_channelPandasIndex
PandasIndex(Index(['xmc/OMV/110', 'xmc/OMV/210', 'xmc/OMV/310'], dtype='object', name='b_field_pol_probe_omv_channel'))
- b_field_pol_probe_omv_geometry_channelPandasIndex
PandasIndex(Index(['omv_201', 'omv_202', 'omv_203', 'omv_204', 'omv_205', 'omv_206', 'omv_207', 'omv_208', 'omv_209', 'omv_210', 'omv_211', 'omv_212', 'omv_213', 'omv_214', 'omv_215', 'omv_216', 'omv_217', 'omv_218', 'omv_219', 'omv_110', 'omv_310'], dtype='object', name='b_field_pol_probe_omv_geometry_channel')) - b_field_tor_probe_cc_channelPandasIndex
PandasIndex(Index(['xmc/CC/MT/201', 'xmc/CC/MT/206', 'xmc/CC/MT/212'], dtype='object', name='b_field_tor_probe_cc_channel'))
- b_field_tor_probe_cc_geometry_channelPandasIndex
PandasIndex(Index(['cc_mt_101', 'cc_mt_102', 'cc_mt_103', 'cc_mt_104', 'cc_mt_105', 'cc_mt_106', 'cc_mt_107', 'cc_mt_108', 'cc_mt_109', 'cc_mt_110', 'cc_mt_111', 'cc_mt_112', 'cc_mt_201', 'cc_mt_202', 'cc_mt_203', 'cc_mt_204', 'cc_mt_205', 'cc_mt_206', 'cc_mt_207', 'cc_mt_208', 'cc_mt_209', 'cc_mt_210', 'cc_mt_211', 'cc_mt_212', 'cc_mt_301', 'cc_mt_302', 'cc_mt_303', 'cc_mt_304', 'cc_mt_305', 'cc_mt_306', 'cc_mt_307', 'cc_mt_308', 'cc_mt_309', 'cc_mt_310', 'cc_mt_311', 'cc_mt_312'], dtype='object', name='b_field_tor_probe_cc_geometry_channel')) - b_field_tor_probe_omaha_channelPandasIndex
PandasIndex(Index(['/xmo/OMAHA/1LZ', '/xmo/OMAHA/3LZ', '/xmo/OMAHA/5LZ', '/xmo/OMAHA/6LZ'], dtype='object', name='b_field_tor_probe_omaha_channel'))
- b_field_tor_probe_saddle_field_channelPandasIndex
PandasIndex(Index(['ASM_SAD/M01', 'ASM_SAD/M02', 'ASM_SAD/M03', 'ASM_SAD/M04', 'ASM_SAD/M05', 'ASM_SAD/M06', 'ASM_SAD/M07', 'ASM_SAD/M08', 'ASM_SAD/M09', 'ASM_SAD/M10', 'ASM_SAD/M11', 'ASM_SAD/M12'], dtype='object', name='b_field_tor_probe_saddle_field_channel')) - b_field_tor_probe_saddle_l_geometry_channelPandasIndex
PandasIndex(Index(['sad_out_l01', 'sad_out_l02', 'sad_out_l03', 'sad_out_l04', 'sad_out_l05', 'sad_out_l06', 'sad_out_l07', 'sad_out_l08', 'sad_out_l09', 'sad_out_l10', 'sad_out_l11', 'sad_out_l12'], dtype='object', name='b_field_tor_probe_saddle_l_geometry_channel')) - b_field_tor_probe_saddle_m_geometry_channelPandasIndex
PandasIndex(Index(['sad_out_m01', 'sad_out_m02', 'sad_out_m03', 'sad_out_m04', 'sad_out_m05', 'sad_out_m06', 'sad_out_m07', 'sad_out_m08', 'sad_out_m09', 'sad_out_m10', 'sad_out_m11', 'sad_out_m12'], dtype='object', name='b_field_tor_probe_saddle_m_geometry_channel')) - b_field_tor_probe_saddle_u_geometry_channelPandasIndex
PandasIndex(Index(['sad_out_u01', 'sad_out_u02', 'sad_out_u03', 'sad_out_u04', 'sad_out_u05', 'sad_out_u06', 'sad_out_u07', 'sad_out_u08', 'sad_out_u09', 'sad_out_u10', 'sad_out_u11', 'sad_out_u12'], dtype='object', name='b_field_tor_probe_saddle_u_geometry_channel')) - b_field_tor_probe_saddle_voltage_channelPandasIndex
PandasIndex(Index(['XMB/SAD/OUT/M01', 'XMB/SAD/OUT/M02', 'XMB/SAD/OUT/M03', 'XMB/SAD/OUT/M04', 'XMB/SAD/OUT/M05', 'XMB/SAD/OUT/M06', 'XMB/SAD/OUT/M07', 'XMB/SAD/OUT/M08', 'XMB/SAD/OUT/M09', 'XMB/SAD/OUT/M10', 'XMB/SAD/OUT/M11', 'XMB/SAD/OUT/M12'], dtype='object', name='b_field_tor_probe_saddle_voltage_channel')) - coordinatePandasIndex
PandasIndex(Index([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27], dtype='int64', name='coordinate')) - flux_loop_channelPandasIndex
PandasIndex(Index(['AMB_FL/CC03', 'AMB_FL/CC04', 'AMB_FL/CC05', 'AMB_FL/CC07', 'AMB_FL/CC09', 'AMB_FL/P3L/4', 'AMB_FL/P3U/1', 'AMB_FL/P3U/4', 'AMB_FL/P4L/1', 'AMB_FL/P4L/4', 'AMB_FL/P4U/4', 'AMB_FL/P5L/1', 'AMB_FL/P5L/4', 'AMB_FL/P5U/1', 'AMB_FL/P6L/1'], dtype='object', name='flux_loop_channel')) - flux_loop_geometry_channelPandasIndex
PandasIndex(Index(['FL_P2U_1', 'FL_P2U_2', 'FL_P2U_3', 'FL_P2U_4', 'FL_P2L_1', 'FL_P2L_2', 'FL_P2L_3', 'FL_P2L_4', 'FL_P3U_1', 'FL_P3U_2', 'FL_P3U_3', 'FL_P3U_4', 'FL_P3L_1', 'FL_P3L_2', 'FL_P3L_3', 'FL_P3L_4', 'FL_P4U_1', 'FL_P4U_2', 'FL_P4U_3', 'FL_P4U_4', 'FL_P4L_1', 'FL_P4L_2', 'FL_P4L_3', 'FL_P4L_4', 'FL_P5U_1', 'FL_P5U_2', 'FL_P5U_3', 'FL_P5U_4', 'FL_P5L_1', 'FL_P5L_2', 'FL_P5L_3', 'FL_P5L_4', 'FL_P6L_1', 'FL_P6L_2', 'FL_CC01', 'FL_CC02', 'FL_CC03', 'FL_CC04', 'FL_CC05', 'FL_CC06', 'FL_CC07', 'FL_CC08', 'FL_CC09', 'FL_CC010'], dtype='object', name='flux_loop_geometry_channel')) - timePandasIndex
PandasIndex(Index([ -0.06120026111602783, -0.06100026111602783, -0.060800261116027834, -0.060600261116027836, -0.06040026111602784, -0.06020026111602784, -0.06000026111602784, -0.05980026111602784, -0.05960026111602784, -0.05940026111602784, ... 0.6633997388839677, 0.6635997388839678, 0.6637997388839678, 0.6639997388839678, 0.6641997388839678, 0.6643997388839677, 0.6645997388839677, 0.6647997388839678, 0.6649997388839678, 0.6651997388839678], dtype='float64', name='time', length=3633)) - time_mirnovPandasIndex
PandasIndex(Index([ -0.06120026111602783, -0.06119826111602783, -0.06119626111602783, -0.061194261116027826, -0.061192261116027824, -0.06119026111602782, -0.06118826111602782, -0.06118626111602782, -0.061184261116027816, -0.061182261116027814, ... 0.6651817388846986, 0.6651837388846986, 0.6651857388846987, 0.6651877388846986, 0.6651897388846986, 0.6651917388846986, 0.6651937388846987, 0.6651957388846986, 0.6651977388846986, 0.6651997388846986], dtype='float64', name='time_mirnov', length=363201)) - time_omahaPandasIndex
PandasIndex(Index([ -0.06120026111602783, -0.06120016111602783, -0.061200061116027826, -0.06119996111602782, -0.06119986111602782, -0.06119976111602782, -0.061199661116027815, -0.06119956111602781, -0.06119946111602781, -0.061199361116027806, ... 0.6651988389048602, 0.6651989389048603, 0.6651990389048603, 0.6651991389048603, 0.6651992389048602, 0.6651993389048603, 0.6651994389048603, 0.6651995389048603, 0.6651996389048602, 0.6651997389048603], dtype='float64', name='time_omaha', length=7264001)) - time_saddlePandasIndex
PandasIndex(Index([ -0.06120026111602783, -0.06118026111602783, -0.061160261116027834, -0.061140261116027834, -0.061120261116027835, -0.061100261116027836, -0.06108026111602784, -0.06106026111602784, -0.06104026111602784, -0.06102026111602784, ... 0.6650197388839426, 0.6650397388839426, 0.6650597388839425, 0.6650797388839426, 0.6650997388839426, 0.6651197388839426, 0.6651397388839426, 0.6651597388839425, 0.6651797388839426, 0.6651997388839426], dtype='float64', name='time_saddle', length=36321))
- name :
- magnetics
- description :
- Magnetic diagnostics for equilibrium identification and plasma shape control.
- imas :
- magnetics
- license_name :
- Creative Commons 4.0 BY-SA
- license_url :
- https://creativecommons.org/licenses/by-sa/4.0/
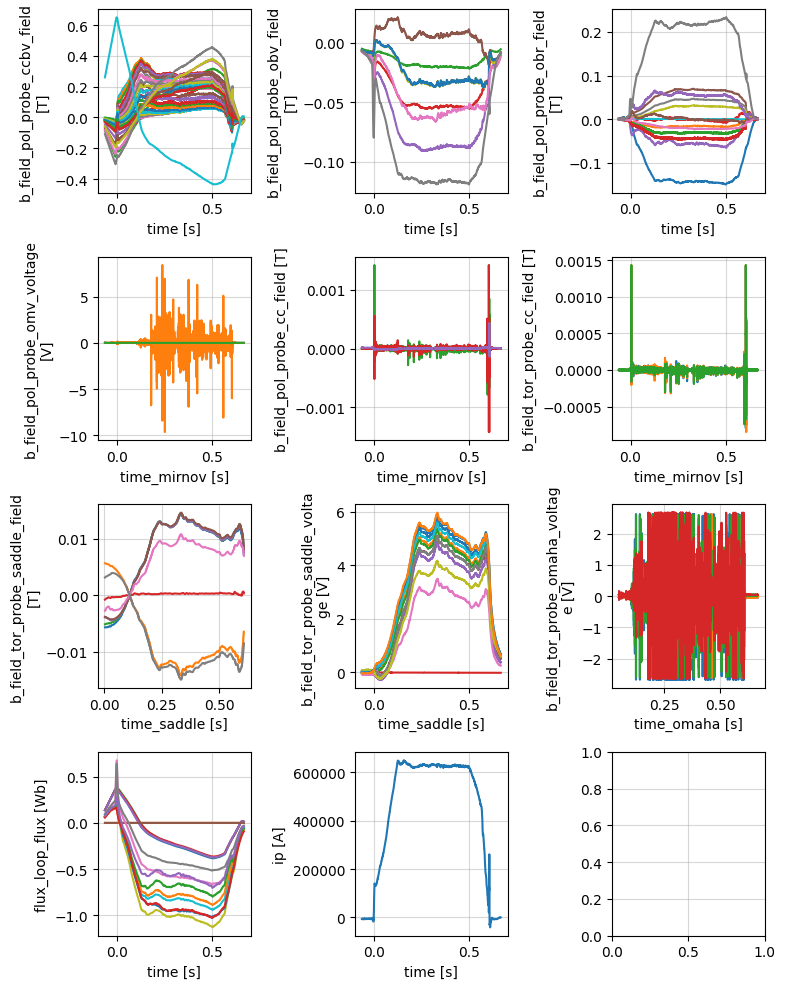
Looking at the spectrogram of one of the mirnov coils can show us information about the MHD modes. Here we see several mode instabilities occuring before the plasma is lost.
ds = profiles['b_field_pol_probe_omv_voltage'].isel(b_field_pol_probe_omv_channel=1)
# Parameters to limit the number of frequencies
nperseg = 2000 # Number of points per segment
nfft = 2000 # Number of FFT points
# Compute the Short-Time Fourier Transform (STFT)
sample_rate = 1/(ds.time_mirnov[1] - ds.time_mirnov[0])
f, t, Zxx = stft(ds, fs=int(sample_rate), nperseg=nperseg, nfft=nfft)
fig, ax = plt.subplots(figsize=(15, 5))
cax = ax.pcolormesh(t, f/1000, np.abs(Zxx), shading='nearest', cmap='jet', norm=LogNorm(vmin=1e-5))
ax.set_ylim(0, 50)
ax.set_title(f'Shot {shot_id}')
ax.set_ylabel('Frequency [Hz]')
ax.set_xlabel('Time [sec]')
plt.colorbar(cax, ax=ax)
plt.show()
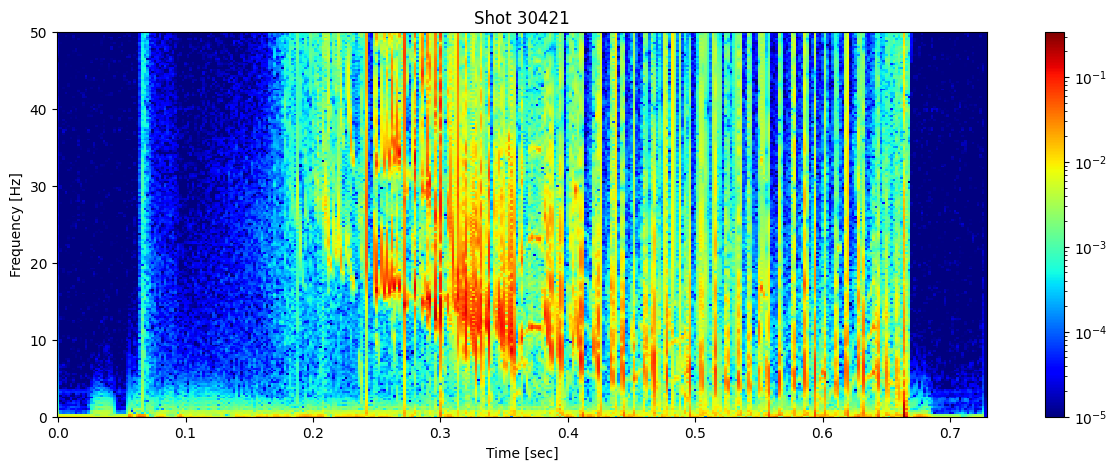
Spectrometer Visible#
Spectrometer in visible light range diagnostic
profiles = xr.open_zarr(store, group='spectrometer_visible')
profiles['filter_spectrometer_dalpha_voltage'].plot.line(x='time')
profiles['filter_spectrometer_bes_voltage'].isel(bes_channel=0).plot.line(x='time_bes')
profiles
<xarray.Dataset> Size: 385MB
Dimensions: (time: 36321, bes_channel: 32,
time_bes: 1452801, dalpha_channel: 3)
Coordinates:
* bes_channel (bes_channel) <U13 2kB 'xbt/channel01...
* dalpha_channel (dalpha_channel) <U13 156B 'XIM_DA/HM...
* time (time) float64 291kB -0.0612 ... 0.6652
* time_bes (time_bes) float64 12MB -0.0612 ... 0...
Data variables:
density_gradient (time) float64 291kB ...
filter_spectrometer_bes_voltage (bes_channel, time_bes) float64 372MB ...
filter_spectrometer_dalpha_voltage (dalpha_channel, time) float64 872kB ...
Attributes:
name: spectrometer_visible
description: Spectrometer in visible light range diagnostic
imas: spectrometer_visible
license_name: Creative Commons 4.0 BY-SA
license_url: https://creativecommons.org/licenses/by-sa/4.0/- time: 36321
- bes_channel: 32
- time_bes: 1452801
- dalpha_channel: 3
- bes_channel(bes_channel)<U13'xbt/channel01' ... 'xbt/channel32'
array(['xbt/channel01', 'xbt/channel02', 'xbt/channel03', 'xbt/channel04', 'xbt/channel05', 'xbt/channel06', 'xbt/channel07', 'xbt/channel08', 'xbt/channel09', 'xbt/channel10', 'xbt/channel11', 'xbt/channel12', 'xbt/channel13', 'xbt/channel14', 'xbt/channel15', 'xbt/channel16', 'xbt/channel17', 'xbt/channel18', 'xbt/channel19', 'xbt/channel20', 'xbt/channel21', 'xbt/channel22', 'xbt/channel23', 'xbt/channel24', 'xbt/channel25', 'xbt/channel26', 'xbt/channel27', 'xbt/channel28', 'xbt/channel29', 'xbt/channel30', 'xbt/channel31', 'xbt/channel32'], dtype='<U13') - dalpha_channel(dalpha_channel)<U13'XIM_DA/HM10/R' ... 'XIM_DA/HU10/T'
array(['XIM_DA/HM10/R', 'XIM_DA/HM10/T', 'XIM_DA/HU10/T'], dtype='<U13')
- time(time)float64-0.0612 -0.06118 ... 0.6652 0.6652
- units :
- s
array([-0.0612 , -0.06118, -0.06116, ..., 0.66516, 0.66518, 0.6652 ], shape=(36321,)) - time_bes(time_bes)float64-0.0612 -0.0612 ... 0.6652 0.6652
- units :
- s
array([-0.0612 , -0.0612 , -0.061199, ..., 0.665199, 0.665199, 0.6652 ], shape=(1452801,))
- density_gradient(time)float64...
- data_block_size :
- 119084
- description :
- label :
- m-4
- name :
- density_gradient
- uda_name :
- ADG_DENSITY_GRADIENT
- units :
- 1 / m ** 4
[36321 values with dtype=float64]
- filter_spectrometer_bes_voltage(bes_channel, time_bes)float64...
- data_block_size :
- 12672876
- description :
- label :
- Volt
- name :
- filter_spectrometer_bes_voltage
- uda_name :
- xbt/channel01
- imas :
- spectrometer_visible.channel[:].filter_spectrometer.photoelectric_voltage.data
- units :
- V
[46489632 values with dtype=float64]
- filter_spectrometer_dalpha_voltage(dalpha_channel, time)float64...
- data_block_size :
- 714428
- description :
- label :
- Volt
- name :
- filter_spectrometer_dalpha_voltage
- uda_name :
- XIM_DA/HM10/R
- imas :
- spectrometer_visible.channel[:].filter_spectrometer.photoelectric_voltage.data
- units :
- V
[108963 values with dtype=float64]
- bes_channelPandasIndex
PandasIndex(Index(['xbt/channel01', 'xbt/channel02', 'xbt/channel03', 'xbt/channel04', 'xbt/channel05', 'xbt/channel06', 'xbt/channel07', 'xbt/channel08', 'xbt/channel09', 'xbt/channel10', 'xbt/channel11', 'xbt/channel12', 'xbt/channel13', 'xbt/channel14', 'xbt/channel15', 'xbt/channel16', 'xbt/channel17', 'xbt/channel18', 'xbt/channel19', 'xbt/channel20', 'xbt/channel21', 'xbt/channel22', 'xbt/channel23', 'xbt/channel24', 'xbt/channel25', 'xbt/channel26', 'xbt/channel27', 'xbt/channel28', 'xbt/channel29', 'xbt/channel30', 'xbt/channel31', 'xbt/channel32'], dtype='object', name='bes_channel')) - dalpha_channelPandasIndex
PandasIndex(Index(['XIM_DA/HM10/R', 'XIM_DA/HM10/T', 'XIM_DA/HU10/T'], dtype='object', name='dalpha_channel'))
- timePandasIndex
PandasIndex(Index([ -0.06120026111602783, -0.06118026111602783, -0.061160261116027834, -0.061140261116027834, -0.061120261116027835, -0.061100261116027836, -0.06108026111602784, -0.06106026111602784, -0.06104026111602784, -0.06102026111602784, ... 0.6650197388839426, 0.6650397388839426, 0.6650597388839425, 0.6650797388839426, 0.6650997388839426, 0.6651197388839426, 0.6651397388839426, 0.6651597388839425, 0.6651797388839426, 0.6651997388839426], dtype='float64', name='time', length=36321)) - time_besPandasIndex
PandasIndex(Index([-0.06120026111602783, -0.06119976111602783, -0.06119926111602783, -0.06119876111602783, -0.06119826111602783, -0.06119776111602783, -0.06119726111602783, -0.06119676111602783, -0.06119626111602783, -0.06119576111602783, ... 0.6651952388846987, 0.6651957388846986, 0.6651962388846986, 0.6651967388846987, 0.6651972388846986, 0.6651977388846986, 0.6651982388846986, 0.6651987388846986, 0.6651992388846987, 0.6651997388846986], dtype='float64', name='time_bes', length=1452801))
- name :
- spectrometer_visible
- description :
- Spectrometer in visible light range diagnostic
- imas :
- spectrometer_visible
- license_name :
- Creative Commons 4.0 BY-SA
- license_url :
- https://creativecommons.org/licenses/by-sa/4.0/
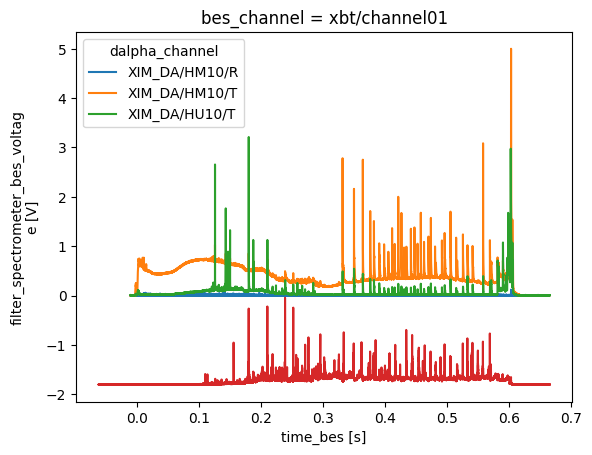
PF Active#
profiles = xr.open_zarr(store, group='pf_active')
fig, axes = plt.subplots(3, 1)
axes = axes.flatten()
profiles['coil_current'].plot.line(x='time', ax=axes[0], add_legend=False)
profiles['coil_voltage'].plot.line(x='time', ax=axes[1], add_legend=False)
profiles['solenoid_current'].plot.line(x='time', ax=axes[2], add_legend=False)
plt.tight_layout()
profiles
<xarray.Dataset> Size: 437kB
Dimensions: (current_channel: 10, time: 2906,
voltage_channel: 4,
p2_inner_lower_coordinate_element: 12,
p2_inner_upper_coordinate_element: 12,
p2_outer_lower_coordinate_element: 8,
p2_outer_upper_coordinate_element: 8,
...
p4_upper_coordinate_element: 23,
p5_lower_coordinate_element: 23,
p5_upper_coordinate_element: 23,
p6_lower_coordinate_element: 4,
p6_upper_coordinate_element: 4,
sol_coordinate_element: 656)
Coordinates: (12/16)
* current_channel (current_channel) <U21 840B 'AMC_P2IL ...
* p2_inner_lower_coordinate_element (p2_inner_lower_coordinate_element) <U15 720B ...
* p2_inner_upper_coordinate_element (p2_inner_upper_coordinate_element) <U15 720B ...
* p2_outer_lower_coordinate_element (p2_outer_lower_coordinate_element) <U14 448B ...
* p2_outer_upper_coordinate_element (p2_outer_upper_coordinate_element) <U14 448B ...
* p3_lower_coordinate_element (p3_lower_coordinate_element) <U14 448B ...
... ...
* p5_upper_coordinate_element (p5_upper_coordinate_element) <U15 1kB ...
* p6_lower_coordinate_element (p6_lower_coordinate_element) <U14 224B ...
* p6_upper_coordinate_element (p6_upper_coordinate_element) <U14 224B ...
* sol_coordinate_element (sol_coordinate_element) <U16 42kB 'co...
* time (time) float64 23kB -0.0612 ... 0.665
* voltage_channel (voltage_channel) <U12 192B '/xdc/pf/f...
Data variables: (12/55)
coil_current (current_channel, time) float64 232kB ...
coil_voltage (voltage_channel, time) float64 93kB ...
p2_inner_lower_height (p2_inner_lower_coordinate_element) float32 48B ...
p2_inner_lower_r (p2_inner_lower_coordinate_element) float32 48B ...
p2_inner_lower_width (p2_inner_lower_coordinate_element) float32 48B ...
p2_inner_lower_z (p2_inner_lower_coordinate_element) float32 48B ...
... ...
p6_upper_z (p6_upper_coordinate_element) float32 16B ...
sol_height (sol_coordinate_element) float32 3kB ...
sol_r (sol_coordinate_element) float32 3kB ...
sol_width (sol_coordinate_element) float32 3kB ...
sol_z (sol_coordinate_element) float32 3kB ...
solenoid_current (time) float64 23kB 8.764e+03 ... 780.2
Attributes:
name: pf_active
description:
imas: pf_active
license_name: Creative Commons 4.0 BY-SA
license_url: https://creativecommons.org/licenses/by-sa/4.0/- current_channel: 10
- time: 2906
- voltage_channel: 4
- p2_inner_lower_coordinate_element: 12
- p2_inner_upper_coordinate_element: 12
- p2_outer_lower_coordinate_element: 8
- p2_outer_upper_coordinate_element: 8
- p3_lower_coordinate_element: 8
- p3_upper_coordinate_element: 8
- p4_lower_coordinate_element: 23
- p4_upper_coordinate_element: 23
- p5_lower_coordinate_element: 23
- p5_upper_coordinate_element: 23
- p6_lower_coordinate_element: 4
- p6_upper_coordinate_element: 4
- sol_coordinate_element: 656
- current_channel(current_channel)<U21'AMC_P2IL FEED CURRENT' ... 'AMC...
array(['AMC_P2IL FEED CURRENT', 'AMC_P2IU FEED CURRENT', 'AMC_P2OL FEED CURRENT', 'AMC_P2OU FEED CURRENT', 'AMC_P3L FEED CURRENT', 'AMC_P3U FEED CURRENT', 'AMC_P4L FEED CURRENT', 'AMC_P4U FEED CURRENT', 'AMC_P5L FEED CURRENT', 'AMC_P5U FEED CURRENT'], dtype='<U21') - p2_inner_lower_coordinate_element(p2_inner_lower_coordinate_element)<U15'coil_element_0' ... 'coil_eleme...
array(['coil_element_0', 'coil_element_1', 'coil_element_2', 'coil_element_3', 'coil_element_4', 'coil_element_5', 'coil_element_6', 'coil_element_7', 'coil_element_8', 'coil_element_9', 'coil_element_10', 'coil_element_11'], dtype='<U15') - p2_inner_upper_coordinate_element(p2_inner_upper_coordinate_element)<U15'coil_element_0' ... 'coil_eleme...
array(['coil_element_0', 'coil_element_1', 'coil_element_2', 'coil_element_3', 'coil_element_4', 'coil_element_5', 'coil_element_6', 'coil_element_7', 'coil_element_8', 'coil_element_9', 'coil_element_10', 'coil_element_11'], dtype='<U15') - p2_outer_lower_coordinate_element(p2_outer_lower_coordinate_element)<U14'coil_element_0' ... 'coil_eleme...
array(['coil_element_0', 'coil_element_1', 'coil_element_2', 'coil_element_3', 'coil_element_4', 'coil_element_5', 'coil_element_6', 'coil_element_7'], dtype='<U14') - p2_outer_upper_coordinate_element(p2_outer_upper_coordinate_element)<U14'coil_element_0' ... 'coil_eleme...
array(['coil_element_0', 'coil_element_1', 'coil_element_2', 'coil_element_3', 'coil_element_4', 'coil_element_5', 'coil_element_6', 'coil_element_7'], dtype='<U14') - p3_lower_coordinate_element(p3_lower_coordinate_element)<U14'coil_element_0' ... 'coil_eleme...
array(['coil_element_0', 'coil_element_1', 'coil_element_2', 'coil_element_3', 'coil_element_4', 'coil_element_5', 'coil_element_6', 'coil_element_7'], dtype='<U14') - p3_upper_coordinate_element(p3_upper_coordinate_element)<U14'coil_element_0' ... 'coil_eleme...
array(['coil_element_0', 'coil_element_1', 'coil_element_2', 'coil_element_3', 'coil_element_4', 'coil_element_5', 'coil_element_6', 'coil_element_7'], dtype='<U14') - p4_lower_coordinate_element(p4_lower_coordinate_element)<U15'coil_element_0' ... 'coil_eleme...
array(['coil_element_0', 'coil_element_1', 'coil_element_2', 'coil_element_3', 'coil_element_4', 'coil_element_5', 'coil_element_6', 'coil_element_7', 'coil_element_8', 'coil_element_9', 'coil_element_10', 'coil_element_11', 'coil_element_12', 'coil_element_13', 'coil_element_14', 'coil_element_15', 'coil_element_16', 'coil_element_17', 'coil_element_18', 'coil_element_19', 'coil_element_20', 'coil_element_21', 'coil_element_22'], dtype='<U15') - p4_upper_coordinate_element(p4_upper_coordinate_element)<U15'coil_element_0' ... 'coil_eleme...
array(['coil_element_0', 'coil_element_1', 'coil_element_2', 'coil_element_3', 'coil_element_4', 'coil_element_5', 'coil_element_6', 'coil_element_7', 'coil_element_8', 'coil_element_9', 'coil_element_10', 'coil_element_11', 'coil_element_12', 'coil_element_13', 'coil_element_14', 'coil_element_15', 'coil_element_16', 'coil_element_17', 'coil_element_18', 'coil_element_19', 'coil_element_20', 'coil_element_21', 'coil_element_22'], dtype='<U15') - p5_lower_coordinate_element(p5_lower_coordinate_element)<U15'coil_element_0' ... 'coil_eleme...
array(['coil_element_0', 'coil_element_1', 'coil_element_2', 'coil_element_3', 'coil_element_4', 'coil_element_5', 'coil_element_6', 'coil_element_7', 'coil_element_8', 'coil_element_9', 'coil_element_10', 'coil_element_11', 'coil_element_12', 'coil_element_13', 'coil_element_14', 'coil_element_15', 'coil_element_16', 'coil_element_17', 'coil_element_18', 'coil_element_19', 'coil_element_20', 'coil_element_21', 'coil_element_22'], dtype='<U15') - p5_upper_coordinate_element(p5_upper_coordinate_element)<U15'coil_element_0' ... 'coil_eleme...
array(['coil_element_0', 'coil_element_1', 'coil_element_2', 'coil_element_3', 'coil_element_4', 'coil_element_5', 'coil_element_6', 'coil_element_7', 'coil_element_8', 'coil_element_9', 'coil_element_10', 'coil_element_11', 'coil_element_12', 'coil_element_13', 'coil_element_14', 'coil_element_15', 'coil_element_16', 'coil_element_17', 'coil_element_18', 'coil_element_19', 'coil_element_20', 'coil_element_21', 'coil_element_22'], dtype='<U15') - p6_lower_coordinate_element(p6_lower_coordinate_element)<U14'coil_element_0' ... 'coil_eleme...
array(['coil_element_0', 'coil_element_1', 'coil_element_2', 'coil_element_3'], dtype='<U14') - p6_upper_coordinate_element(p6_upper_coordinate_element)<U14'coil_element_0' ... 'coil_eleme...
array(['coil_element_0', 'coil_element_1', 'coil_element_2', 'coil_element_3'], dtype='<U14') - sol_coordinate_element(sol_coordinate_element)<U16'coil_element_0' ... 'coil_eleme...
array(['coil_element_0', 'coil_element_1', 'coil_element_2', ..., 'coil_element_653', 'coil_element_654', 'coil_element_655'], shape=(656,), dtype='<U16') - time(time)float64-0.0612 -0.06095 ... 0.6648 0.665
- units :
- s
array([-0.0612 , -0.06095, -0.0607 , ..., 0.66455, 0.6648 , 0.66505], shape=(2906,)) - voltage_channel(voltage_channel)<U12'/xdc/pf/f/p1' ... '/xdc/pf/f/p5'
array(['/xdc/pf/f/p1', '/xdc/pf/f/p2', '/xdc/pf/f/p4', '/xdc/pf/f/p5'], dtype='<U12')
- coil_current(current_channel, time)float64...
- data_block_size :
- 474428
- description :
- label :
- P2il Feed Current
- name :
- coil_current
- uda_name :
- AMC_P2IL FEED CURRENT
- imas :
- pf_active.coil[:].current.data
- units :
- A
[29060 values with dtype=float64]
- coil_voltage(voltage_channel, time)float64...
- data_block_size :
- 450428
- description :
- label :
- /xdc/pf/f/p1
- name :
- coil_voltage
- uda_name :
- /xdc/pf/f/p1
- imas :
- pf_active.coil[:].voltage.data
- units :
- V
[11624 values with dtype=float64]
- p2_inner_lower_height(p2_inner_lower_coordinate_element)float32...
- device :
- MAST
- class_ :
- magnetics
- system :
- pfcoils
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the pfcoils for MAST
- comment :
- Data sourced from EFIT file
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_pf.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-24
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/EFIT_efit_data_pfcoils-3.dat_M4
- imas :
- description :
- Geometric vertical height values of the P2 inner lower coil elements
[12 values with dtype=float32]
- p2_inner_lower_r(p2_inner_lower_coordinate_element)float32...
- device :
- MAST
- class_ :
- magnetics
- system :
- pfcoils
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the pfcoils for MAST
- comment :
- Data sourced from EFIT file
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_pf.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-24
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/EFIT_efit_data_pfcoils-3.dat_M4
- imas :
- description :
- Geometric centre R values of the P2 inner lower coil elements
[12 values with dtype=float32]
- p2_inner_lower_width(p2_inner_lower_coordinate_element)float32...
- device :
- MAST
- class_ :
- magnetics
- system :
- pfcoils
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the pfcoils for MAST
- comment :
- Data sourced from EFIT file
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_pf.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-24
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/EFIT_efit_data_pfcoils-3.dat_M4
- imas :
- description :
- Geometric width values of the P2 inner lower coil elements
[12 values with dtype=float32]
- p2_inner_lower_z(p2_inner_lower_coordinate_element)float32...
- device :
- MAST
- class_ :
- magnetics
- system :
- pfcoils
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the pfcoils for MAST
- comment :
- Data sourced from EFIT file
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_pf.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-24
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/EFIT_efit_data_pfcoils-3.dat_M4
- imas :
- description :
- Geometric centre Z values of the P2 inner lower coil elements
[12 values with dtype=float32]
- p2_inner_upper_height(p2_inner_upper_coordinate_element)float32...
- device :
- MAST
- class_ :
- magnetics
- system :
- pfcoils
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the pfcoils for MAST
- comment :
- Data sourced from EFIT file
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_pf.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-24
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/EFIT_efit_data_pfcoils-3.dat_M4
- imas :
- description :
- Geometric vertical height values of the P2 inner upper coil elements
[12 values with dtype=float32]
- p2_inner_upper_r(p2_inner_upper_coordinate_element)float32...
- device :
- MAST
- class_ :
- magnetics
- system :
- pfcoils
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the pfcoils for MAST
- comment :
- Data sourced from EFIT file
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_pf.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-24
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/EFIT_efit_data_pfcoils-3.dat_M4
- imas :
- description :
- Geometric centre R values of the P2 inner upper coil elements
[12 values with dtype=float32]
- p2_inner_upper_width(p2_inner_upper_coordinate_element)float32...
- device :
- MAST
- class_ :
- magnetics
- system :
- pfcoils
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the pfcoils for MAST
- comment :
- Data sourced from EFIT file
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_pf.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-24
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/EFIT_efit_data_pfcoils-3.dat_M4
- imas :
- description :
- Geometric width values of the P2 inner upper coil elements
[12 values with dtype=float32]
- p2_inner_upper_z(p2_inner_upper_coordinate_element)float32...
- device :
- MAST
- class_ :
- magnetics
- system :
- pfcoils
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the pfcoils for MAST
- comment :
- Data sourced from EFIT file
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_pf.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-24
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/EFIT_efit_data_pfcoils-3.dat_M4
- imas :
- description :
- Geometric centre Z values of the P2 inner upper coil elements
[12 values with dtype=float32]
- p2_outer_lower_height(p2_outer_lower_coordinate_element)float32...
- device :
- MAST
- class_ :
- magnetics
- system :
- pfcoils
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the pfcoils for MAST
- comment :
- Data sourced from EFIT file
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_pf.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-24
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/EFIT_efit_data_pfcoils-3.dat_M4
- imas :
- description :
- Geometric vertical height values of the P2 outer lower coil elements
[8 values with dtype=float32]
- p2_outer_lower_r(p2_outer_lower_coordinate_element)float32...
- device :
- MAST
- class_ :
- magnetics
- system :
- pfcoils
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the pfcoils for MAST
- comment :
- Data sourced from EFIT file
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_pf.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-24
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/EFIT_efit_data_pfcoils-3.dat_M4
- imas :
- description :
- Geometric centre R values of the P2 outer lower coil elements
[8 values with dtype=float32]
- p2_outer_lower_width(p2_outer_lower_coordinate_element)float32...
- device :
- MAST
- class_ :
- magnetics
- system :
- pfcoils
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the pfcoils for MAST
- comment :
- Data sourced from EFIT file
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_pf.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-24
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/EFIT_efit_data_pfcoils-3.dat_M4
- imas :
- description :
- Geometric width values of the P2 outer lower coil elements
[8 values with dtype=float32]
- p2_outer_lower_z(p2_outer_lower_coordinate_element)float32...
- device :
- MAST
- class_ :
- magnetics
- system :
- pfcoils
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the pfcoils for MAST
- comment :
- Data sourced from EFIT file
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_pf.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-24
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/EFIT_efit_data_pfcoils-3.dat_M4
- imas :
- description :
- Geometric centre Z values of the P2 outer lower coil elements
[8 values with dtype=float32]
- p2_outer_upper_height(p2_outer_upper_coordinate_element)float32...
- device :
- MAST
- class_ :
- magnetics
- system :
- pfcoils
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the pfcoils for MAST
- comment :
- Data sourced from EFIT file
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_pf.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-24
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/EFIT_efit_data_pfcoils-3.dat_M4
- imas :
- description :
- Geometric vertical height values of the P2 outer upper coil elements
[8 values with dtype=float32]
- p2_outer_upper_r(p2_outer_upper_coordinate_element)float32...
- device :
- MAST
- class_ :
- magnetics
- system :
- pfcoils
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the pfcoils for MAST
- comment :
- Data sourced from EFIT file
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_pf.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-24
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/EFIT_efit_data_pfcoils-3.dat_M4
- imas :
- description :
- Geometric centre R values of the P2 outer upper coil elements
[8 values with dtype=float32]
- p2_outer_upper_width(p2_outer_upper_coordinate_element)float32...
- device :
- MAST
- class_ :
- magnetics
- system :
- pfcoils
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the pfcoils for MAST
- comment :
- Data sourced from EFIT file
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_pf.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-24
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/EFIT_efit_data_pfcoils-3.dat_M4
- imas :
- description :
- Geometric width values of the P2 outer upper coil elements
[8 values with dtype=float32]
- p2_outer_upper_z(p2_outer_upper_coordinate_element)float32...
- device :
- MAST
- class_ :
- magnetics
- system :
- pfcoils
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the pfcoils for MAST
- comment :
- Data sourced from EFIT file
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_pf.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-24
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/EFIT_efit_data_pfcoils-3.dat_M4
- imas :
- description :
- Geometric centre Z values of the P2 outer upper coil elements
[8 values with dtype=float32]
- p3_lower_height(p3_lower_coordinate_element)float32...
- device :
- MAST
- class_ :
- magnetics
- system :
- pfcoils
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the pfcoils for MAST
- comment :
- Data sourced from EFIT file
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_pf.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-24
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/EFIT_efit_data_pfcoils-3.dat_M4
- imas :
- description :
- Geometric vertical height values of the P3 lower coil elements
[8 values with dtype=float32]
- p3_lower_r(p3_lower_coordinate_element)float32...
- device :
- MAST
- class_ :
- magnetics
- system :
- pfcoils
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the pfcoils for MAST
- comment :
- Data sourced from EFIT file
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_pf.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-24
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/EFIT_efit_data_pfcoils-3.dat_M4
- imas :
- description :
- Geometric centre R values of the P3 lower coil elements
[8 values with dtype=float32]
- p3_lower_width(p3_lower_coordinate_element)float32...
- device :
- MAST
- class_ :
- magnetics
- system :
- pfcoils
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the pfcoils for MAST
- comment :
- Data sourced from EFIT file
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_pf.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-24
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/EFIT_efit_data_pfcoils-3.dat_M4
- imas :
- description :
- Geometric width values of the P3 lower coil elements
[8 values with dtype=float32]
- p3_lower_z(p3_lower_coordinate_element)float32...
- device :
- MAST
- class_ :
- magnetics
- system :
- pfcoils
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the pfcoils for MAST
- comment :
- Data sourced from EFIT file
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_pf.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-24
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/EFIT_efit_data_pfcoils-3.dat_M4
- imas :
- description :
- Geometric centre Z values of the P3 lower coil elements
[8 values with dtype=float32]
- p3_upper_height(p3_upper_coordinate_element)float32...
- device :
- MAST
- class_ :
- magnetics
- system :
- pfcoils
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the pfcoils for MAST
- comment :
- Data sourced from EFIT file
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_pf.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-24
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/EFIT_efit_data_pfcoils-3.dat_M4
- imas :
- description :
- Geometric vertical height values of the P3 upper coil elements
[8 values with dtype=float32]
- p3_upper_r(p3_upper_coordinate_element)float32...
- device :
- MAST
- class_ :
- magnetics
- system :
- pfcoils
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the pfcoils for MAST
- comment :
- Data sourced from EFIT file
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_pf.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-24
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/EFIT_efit_data_pfcoils-3.dat_M4
- imas :
- description :
- Geometric centre R values of the P3 upper coil elements
[8 values with dtype=float32]
- p3_upper_width(p3_upper_coordinate_element)float32...
- device :
- MAST
- class_ :
- magnetics
- system :
- pfcoils
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the pfcoils for MAST
- comment :
- Data sourced from EFIT file
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_pf.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-24
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/EFIT_efit_data_pfcoils-3.dat_M4
- imas :
- description :
- Geometric width values of the P3 upper coil elements
[8 values with dtype=float32]
- p3_upper_z(p3_upper_coordinate_element)float32...
- device :
- MAST
- class_ :
- magnetics
- system :
- pfcoils
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the pfcoils for MAST
- comment :
- Data sourced from EFIT file
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_pf.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-24
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/EFIT_efit_data_pfcoils-3.dat_M4
- imas :
- description :
- Geometric centre Z values of the P3 upper coil elements
[8 values with dtype=float32]
- p4_lower_height(p4_lower_coordinate_element)float32...
- device :
- MAST
- class_ :
- magnetics
- system :
- pfcoils
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the pfcoils for MAST
- comment :
- Data sourced from EFIT file
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_pf.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-24
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/EFIT_efit_data_pfcoils-3.dat_M4
- imas :
- description :
- Geometric vertical height values of the p4 lower coil elements
[23 values with dtype=float32]
- p4_lower_r(p4_lower_coordinate_element)float32...
- device :
- MAST
- class_ :
- magnetics
- system :
- pfcoils
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the pfcoils for MAST
- comment :
- Data sourced from EFIT file
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_pf.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-24
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/EFIT_efit_data_pfcoils-3.dat_M4
- imas :
- description :
- Geometric centre R values of the p4 lower coil elements
[23 values with dtype=float32]
- p4_lower_width(p4_lower_coordinate_element)float32...
- device :
- MAST
- class_ :
- magnetics
- system :
- pfcoils
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the pfcoils for MAST
- comment :
- Data sourced from EFIT file
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_pf.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-24
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/EFIT_efit_data_pfcoils-3.dat_M4
- imas :
- description :
- Geometric width values of the p4 lower coil elements
[23 values with dtype=float32]
- p4_lower_z(p4_lower_coordinate_element)float32...
- device :
- MAST
- class_ :
- magnetics
- system :
- pfcoils
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the pfcoils for MAST
- comment :
- Data sourced from EFIT file
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_pf.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-24
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/EFIT_efit_data_pfcoils-3.dat_M4
- imas :
- description :
- Geometric centre Z values of the p4 lower coil elements
[23 values with dtype=float32]
- p4_upper_height(p4_upper_coordinate_element)float32...
- device :
- MAST
- class_ :
- magnetics
- system :
- pfcoils
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the pfcoils for MAST
- comment :
- Data sourced from EFIT file
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_pf.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-24
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/EFIT_efit_data_pfcoils-3.dat_M4
- imas :
- description :
- Geometric vertical height values of the p4 upper coil elements
[23 values with dtype=float32]
- p4_upper_r(p4_upper_coordinate_element)float32...
- device :
- MAST
- class_ :
- magnetics
- system :
- pfcoils
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the pfcoils for MAST
- comment :
- Data sourced from EFIT file
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_pf.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-24
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/EFIT_efit_data_pfcoils-3.dat_M4
- imas :
- description :
- Geometric centre R values of the p4 upper coil elements
[23 values with dtype=float32]
- p4_upper_width(p4_upper_coordinate_element)float32...
- device :
- MAST
- class_ :
- magnetics
- system :
- pfcoils
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the pfcoils for MAST
- comment :
- Data sourced from EFIT file
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_pf.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-24
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/EFIT_efit_data_pfcoils-3.dat_M4
- imas :
- description :
- Geometric width values of the p4 upper coil elements
[23 values with dtype=float32]
- p4_upper_z(p4_upper_coordinate_element)float32...
- device :
- MAST
- class_ :
- magnetics
- system :
- pfcoils
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the pfcoils for MAST
- comment :
- Data sourced from EFIT file
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_pf.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-24
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/EFIT_efit_data_pfcoils-3.dat_M4
- imas :
- description :
- Geometric centre Z values of the p4 upper coil elements
[23 values with dtype=float32]
- p5_lower_height(p5_lower_coordinate_element)float32...
- device :
- MAST
- class_ :
- magnetics
- system :
- pfcoils
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the pfcoils for MAST
- comment :
- Data sourced from EFIT file
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_pf.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-24
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/EFIT_efit_data_pfcoils-3.dat_M4
- imas :
- description :
- Geometric vertical height values of the p5 lower coil elements
[23 values with dtype=float32]
- p5_lower_r(p5_lower_coordinate_element)float32...
- device :
- MAST
- class_ :
- magnetics
- system :
- pfcoils
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the pfcoils for MAST
- comment :
- Data sourced from EFIT file
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_pf.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-24
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/EFIT_efit_data_pfcoils-3.dat_M4
- imas :
- description :
- Geometric centre R values of the p5 lower coil elements
[23 values with dtype=float32]
- p5_lower_width(p5_lower_coordinate_element)float32...
- device :
- MAST
- class_ :
- magnetics
- system :
- pfcoils
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the pfcoils for MAST
- comment :
- Data sourced from EFIT file
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_pf.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-24
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/EFIT_efit_data_pfcoils-3.dat_M4
- imas :
- description :
- Geometric width values of the p5 lower coil elements
[23 values with dtype=float32]
- p5_lower_z(p5_lower_coordinate_element)float32...
- device :
- MAST
- class_ :
- magnetics
- system :
- pfcoils
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the pfcoils for MAST
- comment :
- Data sourced from EFIT file
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_pf.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-24
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/EFIT_efit_data_pfcoils-3.dat_M4
- imas :
- description :
- Geometric centre Z values of the p5 lower coil elements
[23 values with dtype=float32]
- p5_upper_height(p5_upper_coordinate_element)float32...
- device :
- MAST
- class_ :
- magnetics
- system :
- pfcoils
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the pfcoils for MAST
- comment :
- Data sourced from EFIT file
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_pf.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-24
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/EFIT_efit_data_pfcoils-3.dat_M4
- imas :
- description :
- Geometric vertical height values of the p5 upper coil elements
[23 values with dtype=float32]
- p5_upper_r(p5_upper_coordinate_element)float32...
- device :
- MAST
- class_ :
- magnetics
- system :
- pfcoils
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the pfcoils for MAST
- comment :
- Data sourced from EFIT file
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_pf.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-24
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/EFIT_efit_data_pfcoils-3.dat_M4
- imas :
- description :
- Geometric centre R values of the p5 upper coil elements
[23 values with dtype=float32]
- p5_upper_width(p5_upper_coordinate_element)float32...
- device :
- MAST
- class_ :
- magnetics
- system :
- pfcoils
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the pfcoils for MAST
- comment :
- Data sourced from EFIT file
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_pf.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-24
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/EFIT_efit_data_pfcoils-3.dat_M4
- imas :
- description :
- Geometric width values of the p5 upper coil elements
[23 values with dtype=float32]
- p5_upper_z(p5_upper_coordinate_element)float32...
- device :
- MAST
- class_ :
- magnetics
- system :
- pfcoils
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the pfcoils for MAST
- comment :
- Data sourced from EFIT file
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_pf.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-24
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/EFIT_efit_data_pfcoils-3.dat_M4
- imas :
- description :
- Geometric centre Z values of the p5 upper coil elements
[23 values with dtype=float32]
- p6_lower_height(p6_lower_coordinate_element)float32...
- device :
- MAST
- class_ :
- magnetics
- system :
- pfcoils
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the pfcoils for MAST
- comment :
- Data sourced from EFIT file
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_pf.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-24
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/EFIT_efit_data_pfcoils-3.dat_M4
- imas :
- description :
- Geometric vertical height values of the p6 lower coil elements
[4 values with dtype=float32]
- p6_lower_r(p6_lower_coordinate_element)float32...
- device :
- MAST
- class_ :
- magnetics
- system :
- pfcoils
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the pfcoils for MAST
- comment :
- Data sourced from EFIT file
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_pf.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-24
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/EFIT_efit_data_pfcoils-3.dat_M4
- imas :
- description :
- Geometric centre R values of the p6 lower coil elements
[4 values with dtype=float32]
- p6_lower_width(p6_lower_coordinate_element)float32...
- device :
- MAST
- class_ :
- magnetics
- system :
- pfcoils
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the pfcoils for MAST
- comment :
- Data sourced from EFIT file
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_pf.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-24
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/EFIT_efit_data_pfcoils-3.dat_M4
- imas :
- description :
- Geometric width values of the p6 lower coil elements
[4 values with dtype=float32]
- p6_lower_z(p6_lower_coordinate_element)float32...
- device :
- MAST
- class_ :
- magnetics
- system :
- pfcoils
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the pfcoils for MAST
- comment :
- Data sourced from EFIT file
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_pf.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-24
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/EFIT_efit_data_pfcoils-3.dat_M4
- imas :
- description :
- Geometric centre Z values of the p6 lower coil elements
[4 values with dtype=float32]
- p6_upper_height(p6_upper_coordinate_element)float32...
- device :
- MAST
- class_ :
- magnetics
- system :
- pfcoils
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the pfcoils for MAST
- comment :
- Data sourced from EFIT file
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_pf.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-24
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/EFIT_efit_data_pfcoils-3.dat_M4
- imas :
- description :
- Geometric vertical height values of the p6 upper coil elements
[4 values with dtype=float32]
- p6_upper_r(p6_upper_coordinate_element)float32...
- device :
- MAST
- class_ :
- magnetics
- system :
- pfcoils
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the pfcoils for MAST
- comment :
- Data sourced from EFIT file
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_pf.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-24
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/EFIT_efit_data_pfcoils-3.dat_M4
- imas :
- description :
- Geometric centre R values of the p6 upper coil elements
[4 values with dtype=float32]
- p6_upper_width(p6_upper_coordinate_element)float32...
- device :
- MAST
- class_ :
- magnetics
- system :
- pfcoils
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the pfcoils for MAST
- comment :
- Data sourced from EFIT file
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_pf.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-24
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/EFIT_efit_data_pfcoils-3.dat_M4
- imas :
- description :
- Geometric width values of the p6 upper coil elements
[4 values with dtype=float32]
- p6_upper_z(p6_upper_coordinate_element)float32...
- device :
- MAST
- class_ :
- magnetics
- system :
- pfcoils
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the pfcoils for MAST
- comment :
- Data sourced from EFIT file
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_pf.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-24
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/EFIT_efit_data_pfcoils-3.dat_M4
- imas :
- description :
- Geometric centre Z values of the p6 upper coil elements
[4 values with dtype=float32]
- sol_height(sol_coordinate_element)float32...
- device :
- MAST
- class_ :
- magnetics
- system :
- pfcoils
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the pfcoils for MAST
- comment :
- Data sourced from EFIT file
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_pf.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-24
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/EFIT_efit_data_pfcoils-3.dat_M4
- imas :
- description :
- Geometric vertical height values of the solenoid coil elements
[656 values with dtype=float32]
- sol_r(sol_coordinate_element)float32...
- device :
- MAST
- class_ :
- magnetics
- system :
- pfcoils
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the pfcoils for MAST
- comment :
- Data sourced from EFIT file
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_pf.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-24
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/EFIT_efit_data_pfcoils-3.dat_M4
- imas :
- description :
- Geometric centre R values of the solenoid coil elements
[656 values with dtype=float32]
- sol_width(sol_coordinate_element)float32...
- device :
- MAST
- class_ :
- magnetics
- system :
- pfcoils
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the pfcoils for MAST
- comment :
- Data sourced from EFIT file
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_pf.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-24
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/EFIT_efit_data_pfcoils-3.dat_M4
- imas :
- description :
- Geometric width values of the solenoid coil elements
[656 values with dtype=float32]
- sol_z(sol_coordinate_element)float32...
- device :
- MAST
- class_ :
- magnetics
- system :
- pfcoils
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the pfcoils for MAST
- comment :
- Data sourced from EFIT file
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 0
- revision :
- 0
- status :
- development
- owner :
- MAST Data Systems Team
- signedOffBy :
- lkogan
- creatorCode :
- python create_netcdf_pf.py
- creatorRepo :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-07-24
- createdBy :
- James Hodson (xq0472)
- source :
- https://git.ccfe.ac.uk/MAST-U/mast-geometry/sources/EFIT_efit_data_pfcoils-3.dat_M4
- imas :
- description :
- Geometric centre Z values of the solenoid coil elements
[656 values with dtype=float32]
- solenoid_current(time)float648.764e+03 8.813e+03 ... 773.8 780.2
- data_block_size :
- 474428
- description :
- label :
- Sol Current
- name :
- solenoid_current
- uda_name :
- AMC_SOL CURRENT
- units :
- A
array([8764.074219, 8813.039062, 8868.389648, ..., 778.0495 , 773.791504, 780.17804 ], shape=(2906,))
- current_channelPandasIndex
PandasIndex(Index(['AMC_P2IL FEED CURRENT', 'AMC_P2IU FEED CURRENT', 'AMC_P2OL FEED CURRENT', 'AMC_P2OU FEED CURRENT', 'AMC_P3L FEED CURRENT', 'AMC_P3U FEED CURRENT', 'AMC_P4L FEED CURRENT', 'AMC_P4U FEED CURRENT', 'AMC_P5L FEED CURRENT', 'AMC_P5U FEED CURRENT'], dtype='object', name='current_channel')) - p2_inner_lower_coordinate_elementPandasIndex
PandasIndex(Index(['coil_element_0', 'coil_element_1', 'coil_element_2', 'coil_element_3', 'coil_element_4', 'coil_element_5', 'coil_element_6', 'coil_element_7', 'coil_element_8', 'coil_element_9', 'coil_element_10', 'coil_element_11'], dtype='object', name='p2_inner_lower_coordinate_element')) - p2_inner_upper_coordinate_elementPandasIndex
PandasIndex(Index(['coil_element_0', 'coil_element_1', 'coil_element_2', 'coil_element_3', 'coil_element_4', 'coil_element_5', 'coil_element_6', 'coil_element_7', 'coil_element_8', 'coil_element_9', 'coil_element_10', 'coil_element_11'], dtype='object', name='p2_inner_upper_coordinate_element')) - p2_outer_lower_coordinate_elementPandasIndex
PandasIndex(Index(['coil_element_0', 'coil_element_1', 'coil_element_2', 'coil_element_3', 'coil_element_4', 'coil_element_5', 'coil_element_6', 'coil_element_7'], dtype='object', name='p2_outer_lower_coordinate_element')) - p2_outer_upper_coordinate_elementPandasIndex
PandasIndex(Index(['coil_element_0', 'coil_element_1', 'coil_element_2', 'coil_element_3', 'coil_element_4', 'coil_element_5', 'coil_element_6', 'coil_element_7'], dtype='object', name='p2_outer_upper_coordinate_element')) - p3_lower_coordinate_elementPandasIndex
PandasIndex(Index(['coil_element_0', 'coil_element_1', 'coil_element_2', 'coil_element_3', 'coil_element_4', 'coil_element_5', 'coil_element_6', 'coil_element_7'], dtype='object', name='p3_lower_coordinate_element')) - p3_upper_coordinate_elementPandasIndex
PandasIndex(Index(['coil_element_0', 'coil_element_1', 'coil_element_2', 'coil_element_3', 'coil_element_4', 'coil_element_5', 'coil_element_6', 'coil_element_7'], dtype='object', name='p3_upper_coordinate_element')) - p4_lower_coordinate_elementPandasIndex
PandasIndex(Index(['coil_element_0', 'coil_element_1', 'coil_element_2', 'coil_element_3', 'coil_element_4', 'coil_element_5', 'coil_element_6', 'coil_element_7', 'coil_element_8', 'coil_element_9', 'coil_element_10', 'coil_element_11', 'coil_element_12', 'coil_element_13', 'coil_element_14', 'coil_element_15', 'coil_element_16', 'coil_element_17', 'coil_element_18', 'coil_element_19', 'coil_element_20', 'coil_element_21', 'coil_element_22'], dtype='object', name='p4_lower_coordinate_element')) - p4_upper_coordinate_elementPandasIndex
PandasIndex(Index(['coil_element_0', 'coil_element_1', 'coil_element_2', 'coil_element_3', 'coil_element_4', 'coil_element_5', 'coil_element_6', 'coil_element_7', 'coil_element_8', 'coil_element_9', 'coil_element_10', 'coil_element_11', 'coil_element_12', 'coil_element_13', 'coil_element_14', 'coil_element_15', 'coil_element_16', 'coil_element_17', 'coil_element_18', 'coil_element_19', 'coil_element_20', 'coil_element_21', 'coil_element_22'], dtype='object', name='p4_upper_coordinate_element')) - p5_lower_coordinate_elementPandasIndex
PandasIndex(Index(['coil_element_0', 'coil_element_1', 'coil_element_2', 'coil_element_3', 'coil_element_4', 'coil_element_5', 'coil_element_6', 'coil_element_7', 'coil_element_8', 'coil_element_9', 'coil_element_10', 'coil_element_11', 'coil_element_12', 'coil_element_13', 'coil_element_14', 'coil_element_15', 'coil_element_16', 'coil_element_17', 'coil_element_18', 'coil_element_19', 'coil_element_20', 'coil_element_21', 'coil_element_22'], dtype='object', name='p5_lower_coordinate_element')) - p5_upper_coordinate_elementPandasIndex
PandasIndex(Index(['coil_element_0', 'coil_element_1', 'coil_element_2', 'coil_element_3', 'coil_element_4', 'coil_element_5', 'coil_element_6', 'coil_element_7', 'coil_element_8', 'coil_element_9', 'coil_element_10', 'coil_element_11', 'coil_element_12', 'coil_element_13', 'coil_element_14', 'coil_element_15', 'coil_element_16', 'coil_element_17', 'coil_element_18', 'coil_element_19', 'coil_element_20', 'coil_element_21', 'coil_element_22'], dtype='object', name='p5_upper_coordinate_element')) - p6_lower_coordinate_elementPandasIndex
PandasIndex(Index(['coil_element_0', 'coil_element_1', 'coil_element_2', 'coil_element_3'], dtype='object', name='p6_lower_coordinate_element'))
- p6_upper_coordinate_elementPandasIndex
PandasIndex(Index(['coil_element_0', 'coil_element_1', 'coil_element_2', 'coil_element_3'], dtype='object', name='p6_upper_coordinate_element'))
- sol_coordinate_elementPandasIndex
PandasIndex(Index(['coil_element_0', 'coil_element_1', 'coil_element_2', 'coil_element_3', 'coil_element_4', 'coil_element_5', 'coil_element_6', 'coil_element_7', 'coil_element_8', 'coil_element_9', ... 'coil_element_646', 'coil_element_647', 'coil_element_648', 'coil_element_649', 'coil_element_650', 'coil_element_651', 'coil_element_652', 'coil_element_653', 'coil_element_654', 'coil_element_655'], dtype='object', name='sol_coordinate_element', length=656)) - timePandasIndex
PandasIndex(Index([-0.06120026111602783, -0.06095026111602783, -0.06070026111602783, -0.06045026111602783, -0.06020026111602783, -0.05995026111602783, -0.05970026111602783, -0.05945026111602783, -0.05920026111602783, -0.05895026111602783, ... 0.6627997388839728, 0.6630497388839728, 0.6632997388839728, 0.6635497388839728, 0.6637997388839728, 0.6640497388839728, 0.6642997388839729, 0.6645497388839728, 0.6647997388839728, 0.6650497388839728], dtype='float64', name='time', length=2906)) - voltage_channelPandasIndex
PandasIndex(Index(['/xdc/pf/f/p1', '/xdc/pf/f/p2', '/xdc/pf/f/p4', '/xdc/pf/f/p5'], dtype='object', name='voltage_channel'))
- name :
- pf_active
- description :
- imas :
- pf_active
- license_name :
- Creative Commons 4.0 BY-SA
- license_url :
- https://creativecommons.org/licenses/by-sa/4.0/
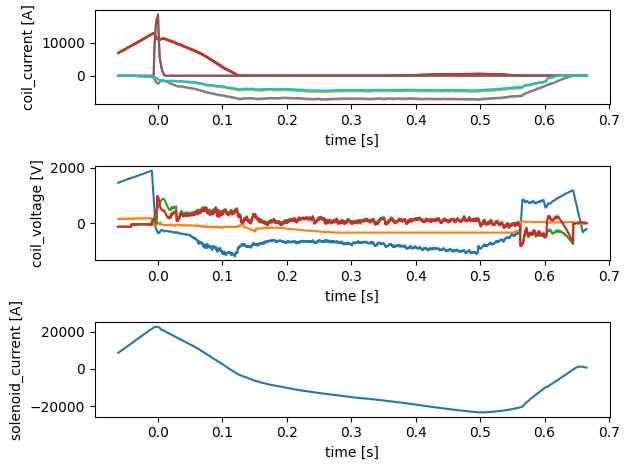
Soft X-rays#
profiles = xr.open_zarr(store, group='soft_x_rays')
fig, axes = plt.subplots(3, 1)
profiles['horizontal_cam_lower'].plot.line(x='time', ax=axes[1], add_legend=False)
axes[1].set_ylim(0, 0.02)
profiles['horizontal_cam_upper'].plot.line(x='time', ax=axes[2], add_legend=False)
axes[2].set_ylim(0, 0.02)
if "tangential_cam" in profiles:
profiles['tangential_cam'].plot.line(x='time', ax=axes[0], add_legend=False)
axes[0].set_ylim(0, 0.2)
plt.tight_layout()
profiles
<xarray.Dataset> Size: 16MB
Dimensions: (horizontal_cam_lower_channel: 18,
time: 36321,
horizontal_cam_lower_geometry_channel: 18,
horizontal_cam_upper_channel: 18,
horizontal_cam_upper_geometry_channel: 18,
inner_vertical_cam_geometry_channel: 12,
outer_vertical_cam_geometry_channel: 12,
tangential_cam_channel: 18,
tangential_cam_geometry_channel: 18,
third_horizontal_cam_geometry_channel: 18)
Coordinates:
* horizontal_cam_lower_channel (horizontal_cam_lower_channel) <U14 1kB ...
* horizontal_cam_lower_geometry_channel (horizontal_cam_lower_geometry_channel) <U23 2kB ...
* horizontal_cam_upper_channel (horizontal_cam_upper_channel) <U14 1kB ...
* horizontal_cam_upper_geometry_channel (horizontal_cam_upper_geometry_channel) <U23 2kB ...
* inner_vertical_cam_geometry_channel (inner_vertical_cam_geometry_channel) <U21 1kB ...
* outer_vertical_cam_geometry_channel (outer_vertical_cam_geometry_channel) <U21 1kB ...
* tangential_cam_channel (tangential_cam_channel) <U12 864B ...
* tangential_cam_geometry_channel (tangential_cam_geometry_channel) <U17 1kB ...
* third_horizontal_cam_geometry_channel (third_horizontal_cam_geometry_channel) <U23 2kB ...
* time (time) float64 291kB -0.0612 ... 0...
Data variables: (12/33)
horizontal_cam_lower (horizontal_cam_lower_channel, time) float64 5MB ...
horizontal_cam_lower_endpoint_r (horizontal_cam_lower_geometry_channel) float64 144B ...
horizontal_cam_lower_endpoint_z (horizontal_cam_lower_geometry_channel) float64 144B ...
horizontal_cam_lower_origin_r (horizontal_cam_lower_geometry_channel) float64 144B ...
horizontal_cam_lower_origin_z (horizontal_cam_lower_geometry_channel) float64 144B ...
horizontal_cam_lower_phi (horizontal_cam_lower_geometry_channel) float64 144B ...
... ...
tangential_cam_phi (tangential_cam_geometry_channel) float64 144B ...
third_horizontal_cam_endpoint_r (third_horizontal_cam_geometry_channel) float64 144B ...
third_horizontal_cam_endpoint_z (third_horizontal_cam_geometry_channel) float64 144B ...
third_horizontal_cam_origin_r (third_horizontal_cam_geometry_channel) float64 144B ...
third_horizontal_cam_origin_z (third_horizontal_cam_geometry_channel) float64 144B ...
third_horizontal_cam_phi (third_horizontal_cam_geometry_channel) float64 144B ...
Attributes:
name: soft_x_rays
description: Soft X-rays tomography diagnostic
imas: soft_x_rays
license_name: Creative Commons 4.0 BY-SA
license_url: https://creativecommons.org/licenses/by-sa/4.0/- horizontal_cam_lower_channel: 18
- time: 36321
- horizontal_cam_lower_geometry_channel: 18
- horizontal_cam_upper_channel: 18
- horizontal_cam_upper_geometry_channel: 18
- inner_vertical_cam_geometry_channel: 12
- outer_vertical_cam_geometry_channel: 12
- tangential_cam_channel: 18
- tangential_cam_geometry_channel: 18
- third_horizontal_cam_geometry_channel: 18
- horizontal_cam_lower_channel(horizontal_cam_lower_channel)<U14'/xsx/HCAM/L/1' ... '/xsx/HCAM/L...
array(['/xsx/HCAM/L/1', '/xsx/HCAM/L/2', '/xsx/HCAM/L/3', '/xsx/HCAM/L/4', '/xsx/HCAM/L/5', '/xsx/HCAM/L/6', '/xsx/HCAM/L/7', '/xsx/HCAM/L/8', '/xsx/HCAM/L/9', '/xsx/HCAM/L/10', '/xsx/HCAM/L/11', '/xsx/HCAM/L/12', '/xsx/HCAM/L/13', '/xsx/HCAM/L/14', '/xsx/HCAM/L/15', '/xsx/HCAM/L/16', '/xsx/HCAM/L/17', '/xsx/HCAM/L/18'], dtype='<U14') - horizontal_cam_lower_geometry_channel(horizontal_cam_lower_geometry_channel)<U23'lower_horizontal_cam_1' ... 'lo...
array(['lower_horizontal_cam_1', 'lower_horizontal_cam_2', 'lower_horizontal_cam_3', 'lower_horizontal_cam_4', 'lower_horizontal_cam_5', 'lower_horizontal_cam_6', 'lower_horizontal_cam_7', 'lower_horizontal_cam_8', 'lower_horizontal_cam_9', 'lower_horizontal_cam_10', 'lower_horizontal_cam_11', 'lower_horizontal_cam_12', 'lower_horizontal_cam_13', 'lower_horizontal_cam_14', 'lower_horizontal_cam_15', 'lower_horizontal_cam_16', 'lower_horizontal_cam_17', 'lower_horizontal_cam_18'], dtype='<U23') - horizontal_cam_upper_channel(horizontal_cam_upper_channel)<U14'/xsx/HCAM/U/1' ... '/xsx/HCAM/U...
array(['/xsx/HCAM/U/1', '/xsx/HCAM/U/2', '/xsx/HCAM/U/3', '/xsx/HCAM/U/4', '/xsx/HCAM/U/5', '/xsx/HCAM/U/6', '/xsx/HCAM/U/7', '/xsx/HCAM/U/8', '/xsx/HCAM/U/9', '/xsx/HCAM/U/10', '/xsx/HCAM/U/11', '/xsx/HCAM/U/12', '/xsx/HCAM/U/13', '/xsx/HCAM/U/14', '/xsx/HCAM/U/15', '/xsx/HCAM/U/16', '/xsx/HCAM/U/17', '/xsx/HCAM/U/18'], dtype='<U14') - horizontal_cam_upper_geometry_channel(horizontal_cam_upper_geometry_channel)<U23'upper_horizontal_cam_1' ... 'up...
array(['upper_horizontal_cam_1', 'upper_horizontal_cam_2', 'upper_horizontal_cam_3', 'upper_horizontal_cam_4', 'upper_horizontal_cam_5', 'upper_horizontal_cam_6', 'upper_horizontal_cam_7', 'upper_horizontal_cam_8', 'upper_horizontal_cam_9', 'upper_horizontal_cam_10', 'upper_horizontal_cam_11', 'upper_horizontal_cam_12', 'upper_horizontal_cam_13', 'upper_horizontal_cam_14', 'upper_horizontal_cam_15', 'upper_horizontal_cam_16', 'upper_horizontal_cam_17', 'upper_horizontal_cam_18'], dtype='<U23') - inner_vertical_cam_geometry_channel(inner_vertical_cam_geometry_channel)<U21'inner_vertical_cam_1' ... 'inne...
array(['inner_vertical_cam_1', 'inner_vertical_cam_2', 'inner_vertical_cam_3', 'inner_vertical_cam_4', 'inner_vertical_cam_5', 'inner_vertical_cam_6', 'inner_vertical_cam_7', 'inner_vertical_cam_8', 'inner_vertical_cam_9', 'inner_vertical_cam_10', 'inner_vertical_cam_11', 'inner_vertical_cam_12'], dtype='<U21') - outer_vertical_cam_geometry_channel(outer_vertical_cam_geometry_channel)<U21'outer_vertical_cam_1' ... 'oute...
array(['outer_vertical_cam_1', 'outer_vertical_cam_2', 'outer_vertical_cam_3', 'outer_vertical_cam_4', 'outer_vertical_cam_5', 'outer_vertical_cam_6', 'outer_vertical_cam_7', 'outer_vertical_cam_8', 'outer_vertical_cam_9', 'outer_vertical_cam_10', 'outer_vertical_cam_11', 'outer_vertical_cam_12'], dtype='<U21') - tangential_cam_channel(tangential_cam_channel)<U12'/xsx/TCAM/1' ... '/xsx/TCAM/18'
array(['/xsx/TCAM/1', '/xsx/TCAM/2', '/xsx/TCAM/3', '/xsx/TCAM/4', '/xsx/TCAM/5', '/xsx/TCAM/6', '/xsx/TCAM/7', '/xsx/TCAM/8', '/xsx/TCAM/9', '/xsx/TCAM/10', '/xsx/TCAM/11', '/xsx/TCAM/12', '/xsx/TCAM/13', '/xsx/TCAM/14', '/xsx/TCAM/15', '/xsx/TCAM/16', '/xsx/TCAM/17', '/xsx/TCAM/18'], dtype='<U12') - tangential_cam_geometry_channel(tangential_cam_geometry_channel)<U17'tangential_cam_1' ... 'tangenti...
array(['tangential_cam_1', 'tangential_cam_2', 'tangential_cam_3', 'tangential_cam_4', 'tangential_cam_5', 'tangential_cam_6', 'tangential_cam_7', 'tangential_cam_8', 'tangential_cam_9', 'tangential_cam_10', 'tangential_cam_11', 'tangential_cam_12', 'tangential_cam_13', 'tangential_cam_14', 'tangential_cam_15', 'tangential_cam_16', 'tangential_cam_17', 'tangential_cam_18'], dtype='<U17') - third_horizontal_cam_geometry_channel(third_horizontal_cam_geometry_channel)<U23'third_horizontal_cam_1' ... 'th...
array(['third_horizontal_cam_1', 'third_horizontal_cam_2', 'third_horizontal_cam_3', 'third_horizontal_cam_4', 'third_horizontal_cam_5', 'third_horizontal_cam_6', 'third_horizontal_cam_7', 'third_horizontal_cam_8', 'third_horizontal_cam_9', 'third_horizontal_cam_10', 'third_horizontal_cam_11', 'third_horizontal_cam_12', 'third_horizontal_cam_13', 'third_horizontal_cam_14', 'third_horizontal_cam_15', 'third_horizontal_cam_16', 'third_horizontal_cam_17', 'third_horizontal_cam_18'], dtype='<U23') - time(time)float64-0.0612 -0.06118 ... 0.6652 0.6652
- units :
- s
array([-0.0612 , -0.06118, -0.06116, ..., 0.66516, 0.66518, 0.6652 ], shape=(36321,))
- horizontal_cam_lower(horizontal_cam_lower_channel, time)float64...
- data_block_size :
- 2514444
- description :
- label :
- Volt
- name :
- horizontal_cam_lower
- uda_name :
- /xsx/HCAM/L/1
- units :
- V
[653778 values with dtype=float64]
- horizontal_cam_lower_endpoint_r(horizontal_cam_lower_geometry_channel)float64...
- device :
- MAST
- class_ :
- xraycams
- system :
- xraycams
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the x-ray cameras for MAST
- comment :
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 1
- revision :
- 0
- status :
- development
- owner :
- jhodson
- signedOffBy :
- creatorCode :
- python create_netcdf_xraycams.py
- creatorRepo :
- MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-05-16
- createdBy :
- sfrankel
- source :
- MAST-U/MAST Geometry/sources/ssx_spec.pdf
- imas :
- description :
- Endpoint line of sight major radius of the lower horizontal camera
[18 values with dtype=float64]
- horizontal_cam_lower_endpoint_z(horizontal_cam_lower_geometry_channel)float64...
- device :
- MAST
- class_ :
- xraycams
- system :
- xraycams
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the x-ray cameras for MAST
- comment :
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 1
- revision :
- 0
- status :
- development
- owner :
- jhodson
- signedOffBy :
- creatorCode :
- python create_netcdf_xraycams.py
- creatorRepo :
- MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-05-16
- createdBy :
- sfrankel
- source :
- MAST-U/MAST Geometry/sources/ssx_spec.pdf
- imas :
- description :
- Endpoint line of sight z of the lower horizontal camera
[18 values with dtype=float64]
- horizontal_cam_lower_origin_r(horizontal_cam_lower_geometry_channel)float64...
- device :
- MAST
- class_ :
- xraycams
- system :
- xraycams
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the x-ray cameras for MAST
- comment :
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 1
- revision :
- 0
- status :
- development
- owner :
- jhodson
- signedOffBy :
- creatorCode :
- python create_netcdf_xraycams.py
- creatorRepo :
- MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-05-16
- createdBy :
- sfrankel
- source :
- MAST-U/MAST Geometry/sources/ssx_spec.pdf
- imas :
- description :
- Origin line of sight radius of the lower horizontal camera
[18 values with dtype=float64]
- horizontal_cam_lower_origin_z(horizontal_cam_lower_geometry_channel)float64...
- device :
- MAST
- class_ :
- xraycams
- system :
- xraycams
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the x-ray cameras for MAST
- comment :
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 1
- revision :
- 0
- status :
- development
- owner :
- jhodson
- signedOffBy :
- creatorCode :
- python create_netcdf_xraycams.py
- creatorRepo :
- MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-05-16
- createdBy :
- sfrankel
- source :
- MAST-U/MAST Geometry/sources/ssx_spec.pdf
- imas :
- description :
- Origin line of sight z of the lower horizontal camera
[18 values with dtype=float64]
- horizontal_cam_lower_phi(horizontal_cam_lower_geometry_channel)float64...
- device :
- MAST
- class_ :
- xraycams
- system :
- xraycams
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the x-ray cameras for MAST
- comment :
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 1
- revision :
- 0
- status :
- development
- owner :
- jhodson
- signedOffBy :
- creatorCode :
- python create_netcdf_xraycams.py
- creatorRepo :
- MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-05-16
- createdBy :
- sfrankel
- source :
- MAST-U/MAST Geometry/sources/ssx_spec.pdf
- imas :
- description :
- Toroidal angle of the lower horizontal camera
[18 values with dtype=float64]
- horizontal_cam_upper(horizontal_cam_upper_channel, time)float64...
- data_block_size :
- 2514444
- description :
- label :
- Volt
- name :
- horizontal_cam_upper
- uda_name :
- /xsx/HCAM/U/1
- units :
- V
[653778 values with dtype=float64]
- horizontal_cam_upper_endpoint_r(horizontal_cam_upper_geometry_channel)float64...
- device :
- MAST
- class_ :
- xraycams
- system :
- xraycams
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the x-ray cameras for MAST
- comment :
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 1
- revision :
- 0
- status :
- development
- owner :
- jhodson
- signedOffBy :
- creatorCode :
- python create_netcdf_xraycams.py
- creatorRepo :
- MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-05-16
- createdBy :
- sfrankel
- source :
- MAST-U/MAST Geometry/sources/ssx_spec.pdf
- imas :
- description :
- Endpoint line of sight major radius of the upper horizontal camera
[18 values with dtype=float64]
- horizontal_cam_upper_endpoint_z(horizontal_cam_upper_geometry_channel)float64...
- device :
- MAST
- class_ :
- xraycams
- system :
- xraycams
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the x-ray cameras for MAST
- comment :
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 1
- revision :
- 0
- status :
- development
- owner :
- jhodson
- signedOffBy :
- creatorCode :
- python create_netcdf_xraycams.py
- creatorRepo :
- MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-05-16
- createdBy :
- sfrankel
- source :
- MAST-U/MAST Geometry/sources/ssx_spec.pdf
- imas :
- description :
- Endpoint line of sight z of the upper horizontal camera
[18 values with dtype=float64]
- horizontal_cam_upper_origin_r(horizontal_cam_upper_geometry_channel)float64...
- device :
- MAST
- class_ :
- xraycams
- system :
- xraycams
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the x-ray cameras for MAST
- comment :
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 1
- revision :
- 0
- status :
- development
- owner :
- jhodson
- signedOffBy :
- creatorCode :
- python create_netcdf_xraycams.py
- creatorRepo :
- MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-05-16
- createdBy :
- sfrankel
- source :
- MAST-U/MAST Geometry/sources/ssx_spec.pdf
- imas :
- description :
- Origin line of sight radius of the upper horizontal camera
[18 values with dtype=float64]
- horizontal_cam_upper_origin_z(horizontal_cam_upper_geometry_channel)float64...
- device :
- MAST
- class_ :
- xraycams
- system :
- xraycams
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the x-ray cameras for MAST
- comment :
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 1
- revision :
- 0
- status :
- development
- owner :
- jhodson
- signedOffBy :
- creatorCode :
- python create_netcdf_xraycams.py
- creatorRepo :
- MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-05-16
- createdBy :
- sfrankel
- source :
- MAST-U/MAST Geometry/sources/ssx_spec.pdf
- imas :
- description :
- Origin line of sight z of the upper horizontal camera
[18 values with dtype=float64]
- horizontal_cam_upper_phi(horizontal_cam_upper_geometry_channel)float64...
- device :
- MAST
- class_ :
- xraycams
- system :
- xraycams
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the x-ray cameras for MAST
- comment :
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 1
- revision :
- 0
- status :
- development
- owner :
- jhodson
- signedOffBy :
- creatorCode :
- python create_netcdf_xraycams.py
- creatorRepo :
- MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-05-16
- createdBy :
- sfrankel
- source :
- MAST-U/MAST Geometry/sources/ssx_spec.pdf
- imas :
- description :
- Toroidal angle of the upper horizontal camera
[18 values with dtype=float64]
- inner_vertical_cam_endpoint_r(inner_vertical_cam_geometry_channel)float64...
- device :
- MAST
- class_ :
- xraycams
- system :
- xraycams
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the x-ray cameras for MAST
- comment :
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 1
- revision :
- 0
- status :
- development
- owner :
- jhodson
- signedOffBy :
- creatorCode :
- python create_netcdf_xraycams.py
- creatorRepo :
- MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-05-16
- createdBy :
- sfrankel
- source :
- MAST-U/MAST Geometry/sources/ssx_spec.pdf
- imas :
- description :
- Endpoint line of sight radius of the inner vertical camera
[12 values with dtype=float64]
- inner_vertical_cam_endpoint_z(inner_vertical_cam_geometry_channel)float64...
- device :
- MAST
- class_ :
- xraycams
- system :
- xraycams
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the x-ray cameras for MAST
- comment :
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 1
- revision :
- 0
- status :
- development
- owner :
- jhodson
- signedOffBy :
- creatorCode :
- python create_netcdf_xraycams.py
- creatorRepo :
- MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-05-16
- createdBy :
- sfrankel
- source :
- MAST-U/MAST Geometry/sources/ssx_spec.pdf
- imas :
- description :
- Endpoint line of sight Z of the inner vertical camera
[12 values with dtype=float64]
- inner_vertical_cam_origin_r(inner_vertical_cam_geometry_channel)float64...
- device :
- MAST
- class_ :
- xraycams
- system :
- xraycams
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the x-ray cameras for MAST
- comment :
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 1
- revision :
- 0
- status :
- development
- owner :
- jhodson
- signedOffBy :
- creatorCode :
- python create_netcdf_xraycams.py
- creatorRepo :
- MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-05-16
- createdBy :
- sfrankel
- source :
- MAST-U/MAST Geometry/sources/ssx_spec.pdf
- imas :
- description :
- Origin line of sight radius of the inner vertical camera
[12 values with dtype=float64]
- inner_vertical_cam_origin_z(inner_vertical_cam_geometry_channel)float64...
- device :
- MAST
- class_ :
- xraycams
- system :
- xraycams
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the x-ray cameras for MAST
- comment :
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 1
- revision :
- 0
- status :
- development
- owner :
- jhodson
- signedOffBy :
- creatorCode :
- python create_netcdf_xraycams.py
- creatorRepo :
- MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-05-16
- createdBy :
- sfrankel
- source :
- MAST-U/MAST Geometry/sources/ssx_spec.pdf
- imas :
- description :
- Origin line of sight Z of the inner vertical camera
[12 values with dtype=float64]
- inner_vertical_cam_phi(inner_vertical_cam_geometry_channel)float64...
- device :
- MAST
- class_ :
- xraycams
- system :
- xraycams
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the x-ray cameras for MAST
- comment :
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 1
- revision :
- 0
- status :
- development
- owner :
- jhodson
- signedOffBy :
- creatorCode :
- python create_netcdf_xraycams.py
- creatorRepo :
- MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-05-16
- createdBy :
- sfrankel
- source :
- MAST-U/MAST Geometry/sources/ssx_spec.pdf
- imas :
- description :
- Toroidal angle of the inner vertical camera
[12 values with dtype=float64]
- outer_vertical_cam_endpoint_r(outer_vertical_cam_geometry_channel)float64...
- device :
- MAST
- class_ :
- xraycams
- system :
- xraycams
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the x-ray cameras for MAST
- comment :
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 1
- revision :
- 0
- status :
- development
- owner :
- jhodson
- signedOffBy :
- creatorCode :
- python create_netcdf_xraycams.py
- creatorRepo :
- MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-05-16
- createdBy :
- sfrankel
- source :
- MAST-U/MAST Geometry/sources/ssx_spec.pdf
- imas :
- description :
- Endpoint line of sight radius of the outer vertical camera
[12 values with dtype=float64]
- outer_vertical_cam_endpoint_z(outer_vertical_cam_geometry_channel)float64...
- device :
- MAST
- class_ :
- xraycams
- system :
- xraycams
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the x-ray cameras for MAST
- comment :
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 1
- revision :
- 0
- status :
- development
- owner :
- jhodson
- signedOffBy :
- creatorCode :
- python create_netcdf_xraycams.py
- creatorRepo :
- MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-05-16
- createdBy :
- sfrankel
- source :
- MAST-U/MAST Geometry/sources/ssx_spec.pdf
- imas :
- description :
- Endpoint line of sight Z of the outer vertical camera
[12 values with dtype=float64]
- outer_vertical_cam_origin_r(outer_vertical_cam_geometry_channel)float64...
- device :
- MAST
- class_ :
- xraycams
- system :
- xraycams
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the x-ray cameras for MAST
- comment :
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 1
- revision :
- 0
- status :
- development
- owner :
- jhodson
- signedOffBy :
- creatorCode :
- python create_netcdf_xraycams.py
- creatorRepo :
- MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-05-16
- createdBy :
- sfrankel
- source :
- MAST-U/MAST Geometry/sources/ssx_spec.pdf
- imas :
- description :
- Origin line of sight radius of the outer vertical camera
[12 values with dtype=float64]
- outer_vertical_cam_origin_z(outer_vertical_cam_geometry_channel)float64...
- device :
- MAST
- class_ :
- xraycams
- system :
- xraycams
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the x-ray cameras for MAST
- comment :
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 1
- revision :
- 0
- status :
- development
- owner :
- jhodson
- signedOffBy :
- creatorCode :
- python create_netcdf_xraycams.py
- creatorRepo :
- MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-05-16
- createdBy :
- sfrankel
- source :
- MAST-U/MAST Geometry/sources/ssx_spec.pdf
- imas :
- description :
- Origin line of sight Z of the outer vertical camera
[12 values with dtype=float64]
- outer_vertical_cam_phi(outer_vertical_cam_geometry_channel)float64...
- device :
- MAST
- class_ :
- xraycams
- system :
- xraycams
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the x-ray cameras for MAST
- comment :
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 1
- revision :
- 0
- status :
- development
- owner :
- jhodson
- signedOffBy :
- creatorCode :
- python create_netcdf_xraycams.py
- creatorRepo :
- MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-05-16
- createdBy :
- sfrankel
- source :
- MAST-U/MAST Geometry/sources/ssx_spec.pdf
- imas :
- description :
- Toroidal angle of the outer vertical camera
[12 values with dtype=float64]
- tangential_cam(tangential_cam_channel, time)float64...
- data_block_size :
- 2514444
- description :
- label :
- Volt
- name :
- tangential_cam
- uda_name :
- /xsx/TCAM/1
- units :
- V
[653778 values with dtype=float64]
- tangential_cam_endpoint_r(tangential_cam_geometry_channel)float64...
- device :
- MAST
- class_ :
- xraycams
- system :
- xraycams
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the x-ray cameras for MAST
- comment :
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 1
- revision :
- 0
- status :
- development
- owner :
- jhodson
- signedOffBy :
- creatorCode :
- python create_netcdf_xraycams.py
- creatorRepo :
- MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-05-16
- createdBy :
- sfrankel
- source :
- MAST-U/MAST Geometry/sources/ssx_spec.pdf
- imas :
- description :
- Endpoint line of sight radius of the tangential camera
[18 values with dtype=float64]
- tangential_cam_endpoint_z(tangential_cam_geometry_channel)float64...
- device :
- MAST
- class_ :
- xraycams
- system :
- xraycams
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the x-ray cameras for MAST
- comment :
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 1
- revision :
- 0
- status :
- development
- owner :
- jhodson
- signedOffBy :
- creatorCode :
- python create_netcdf_xraycams.py
- creatorRepo :
- MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-05-16
- createdBy :
- sfrankel
- source :
- MAST-U/MAST Geometry/sources/ssx_spec.pdf
- imas :
- description :
- Endpoint line of sight z of the tangential camera
[18 values with dtype=float64]
- tangential_cam_origin_r(tangential_cam_geometry_channel)float64...
- device :
- MAST
- class_ :
- xraycams
- system :
- xraycams
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the x-ray cameras for MAST
- comment :
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 1
- revision :
- 0
- status :
- development
- owner :
- jhodson
- signedOffBy :
- creatorCode :
- python create_netcdf_xraycams.py
- creatorRepo :
- MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-05-16
- createdBy :
- sfrankel
- source :
- MAST-U/MAST Geometry/sources/ssx_spec.pdf
- imas :
- description :
- Origin line of sight radius of the tangential camera
[18 values with dtype=float64]
- tangential_cam_origin_z(tangential_cam_geometry_channel)float64...
- device :
- MAST
- class_ :
- xraycams
- system :
- xraycams
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the x-ray cameras for MAST
- comment :
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 1
- revision :
- 0
- status :
- development
- owner :
- jhodson
- signedOffBy :
- creatorCode :
- python create_netcdf_xraycams.py
- creatorRepo :
- MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-05-16
- createdBy :
- sfrankel
- source :
- MAST-U/MAST Geometry/sources/ssx_spec.pdf
- imas :
- description :
- Origin line of sight z of the tangential camera
[18 values with dtype=float64]
- tangential_cam_phi(tangential_cam_geometry_channel)float64...
- device :
- MAST
- class_ :
- xraycams
- system :
- xraycams
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the x-ray cameras for MAST
- comment :
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 1
- revision :
- 0
- status :
- development
- owner :
- jhodson
- signedOffBy :
- creatorCode :
- python create_netcdf_xraycams.py
- creatorRepo :
- MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-05-16
- createdBy :
- sfrankel
- source :
- MAST-U/MAST Geometry/sources/ssx_spec.pdf
- imas :
- description :
- Toroidal angle of the tangential camera
[18 values with dtype=float64]
- third_horizontal_cam_endpoint_r(third_horizontal_cam_geometry_channel)float64...
- device :
- MAST
- class_ :
- xraycams
- system :
- xraycams
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the x-ray cameras for MAST
- comment :
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 1
- revision :
- 0
- status :
- development
- owner :
- jhodson
- signedOffBy :
- creatorCode :
- python create_netcdf_xraycams.py
- creatorRepo :
- MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-05-16
- createdBy :
- sfrankel
- source :
- MAST-U/MAST Geometry/sources/ssx_spec.pdf
- imas :
- description :
- Endpoint line of sight radius of the third horizontal camera
[18 values with dtype=float64]
- third_horizontal_cam_endpoint_z(third_horizontal_cam_geometry_channel)float64...
- device :
- MAST
- class_ :
- xraycams
- system :
- xraycams
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the x-ray cameras for MAST
- comment :
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 1
- revision :
- 0
- status :
- development
- owner :
- jhodson
- signedOffBy :
- creatorCode :
- python create_netcdf_xraycams.py
- creatorRepo :
- MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-05-16
- createdBy :
- sfrankel
- source :
- MAST-U/MAST Geometry/sources/ssx_spec.pdf
- imas :
- description :
- Endpoint line of sight Z of the third horizontal camera
[18 values with dtype=float64]
- third_horizontal_cam_origin_r(third_horizontal_cam_geometry_channel)float64...
- device :
- MAST
- class_ :
- xraycams
- system :
- xraycams
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the x-ray cameras for MAST
- comment :
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 1
- revision :
- 0
- status :
- development
- owner :
- jhodson
- signedOffBy :
- creatorCode :
- python create_netcdf_xraycams.py
- creatorRepo :
- MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-05-16
- createdBy :
- sfrankel
- source :
- MAST-U/MAST Geometry/sources/ssx_spec.pdf
- imas :
- description :
- Origin line of sight radius of the third horizontal camera
[18 values with dtype=float64]
- third_horizontal_cam_origin_z(third_horizontal_cam_geometry_channel)float64...
- device :
- MAST
- class_ :
- xraycams
- system :
- xraycams
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the x-ray cameras for MAST
- comment :
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 1
- revision :
- 0
- status :
- development
- owner :
- jhodson
- signedOffBy :
- creatorCode :
- python create_netcdf_xraycams.py
- creatorRepo :
- MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-05-16
- createdBy :
- sfrankel
- source :
- MAST-U/MAST Geometry/sources/ssx_spec.pdf
- imas :
- description :
- Origin line of sight Z of the third horizontal camera
[18 values with dtype=float64]
- third_horizontal_cam_phi(third_horizontal_cam_geometry_channel)float64...
- device :
- MAST
- class_ :
- xraycams
- system :
- xraycams
- configuration :
- geometry
- shotRangeStart :
- 0
- shotRangeStop :
- 40000
- content :
- geometry of the x-ray cameras for MAST
- comment :
- units :
- SI, degrees, m
- coordinateSystem :
- Cylindrical
- calibration :
- None
- version :
- 1
- revision :
- 0
- status :
- development
- owner :
- jhodson
- signedOffBy :
- creatorCode :
- python create_netcdf_xraycams.py
- creatorRepo :
- MAST-U/mast-geometry
- creatorCommitId :
- creationDate :
- 2025-05-16
- createdBy :
- sfrankel
- source :
- MAST-U/MAST Geometry/sources/ssx_spec.pdf
- imas :
- description :
- Toroidal angle of the third horizontal camera
[18 values with dtype=float64]
- horizontal_cam_lower_channelPandasIndex
PandasIndex(Index(['/xsx/HCAM/L/1', '/xsx/HCAM/L/2', '/xsx/HCAM/L/3', '/xsx/HCAM/L/4', '/xsx/HCAM/L/5', '/xsx/HCAM/L/6', '/xsx/HCAM/L/7', '/xsx/HCAM/L/8', '/xsx/HCAM/L/9', '/xsx/HCAM/L/10', '/xsx/HCAM/L/11', '/xsx/HCAM/L/12', '/xsx/HCAM/L/13', '/xsx/HCAM/L/14', '/xsx/HCAM/L/15', '/xsx/HCAM/L/16', '/xsx/HCAM/L/17', '/xsx/HCAM/L/18'], dtype='object', name='horizontal_cam_lower_channel')) - horizontal_cam_lower_geometry_channelPandasIndex
PandasIndex(Index(['lower_horizontal_cam_1', 'lower_horizontal_cam_2', 'lower_horizontal_cam_3', 'lower_horizontal_cam_4', 'lower_horizontal_cam_5', 'lower_horizontal_cam_6', 'lower_horizontal_cam_7', 'lower_horizontal_cam_8', 'lower_horizontal_cam_9', 'lower_horizontal_cam_10', 'lower_horizontal_cam_11', 'lower_horizontal_cam_12', 'lower_horizontal_cam_13', 'lower_horizontal_cam_14', 'lower_horizontal_cam_15', 'lower_horizontal_cam_16', 'lower_horizontal_cam_17', 'lower_horizontal_cam_18'], dtype='object', name='horizontal_cam_lower_geometry_channel')) - horizontal_cam_upper_channelPandasIndex
PandasIndex(Index(['/xsx/HCAM/U/1', '/xsx/HCAM/U/2', '/xsx/HCAM/U/3', '/xsx/HCAM/U/4', '/xsx/HCAM/U/5', '/xsx/HCAM/U/6', '/xsx/HCAM/U/7', '/xsx/HCAM/U/8', '/xsx/HCAM/U/9', '/xsx/HCAM/U/10', '/xsx/HCAM/U/11', '/xsx/HCAM/U/12', '/xsx/HCAM/U/13', '/xsx/HCAM/U/14', '/xsx/HCAM/U/15', '/xsx/HCAM/U/16', '/xsx/HCAM/U/17', '/xsx/HCAM/U/18'], dtype='object', name='horizontal_cam_upper_channel')) - horizontal_cam_upper_geometry_channelPandasIndex
PandasIndex(Index(['upper_horizontal_cam_1', 'upper_horizontal_cam_2', 'upper_horizontal_cam_3', 'upper_horizontal_cam_4', 'upper_horizontal_cam_5', 'upper_horizontal_cam_6', 'upper_horizontal_cam_7', 'upper_horizontal_cam_8', 'upper_horizontal_cam_9', 'upper_horizontal_cam_10', 'upper_horizontal_cam_11', 'upper_horizontal_cam_12', 'upper_horizontal_cam_13', 'upper_horizontal_cam_14', 'upper_horizontal_cam_15', 'upper_horizontal_cam_16', 'upper_horizontal_cam_17', 'upper_horizontal_cam_18'], dtype='object', name='horizontal_cam_upper_geometry_channel')) - inner_vertical_cam_geometry_channelPandasIndex
PandasIndex(Index(['inner_vertical_cam_1', 'inner_vertical_cam_2', 'inner_vertical_cam_3', 'inner_vertical_cam_4', 'inner_vertical_cam_5', 'inner_vertical_cam_6', 'inner_vertical_cam_7', 'inner_vertical_cam_8', 'inner_vertical_cam_9', 'inner_vertical_cam_10', 'inner_vertical_cam_11', 'inner_vertical_cam_12'], dtype='object', name='inner_vertical_cam_geometry_channel')) - outer_vertical_cam_geometry_channelPandasIndex
PandasIndex(Index(['outer_vertical_cam_1', 'outer_vertical_cam_2', 'outer_vertical_cam_3', 'outer_vertical_cam_4', 'outer_vertical_cam_5', 'outer_vertical_cam_6', 'outer_vertical_cam_7', 'outer_vertical_cam_8', 'outer_vertical_cam_9', 'outer_vertical_cam_10', 'outer_vertical_cam_11', 'outer_vertical_cam_12'], dtype='object', name='outer_vertical_cam_geometry_channel')) - tangential_cam_channelPandasIndex
PandasIndex(Index(['/xsx/TCAM/1', '/xsx/TCAM/2', '/xsx/TCAM/3', '/xsx/TCAM/4', '/xsx/TCAM/5', '/xsx/TCAM/6', '/xsx/TCAM/7', '/xsx/TCAM/8', '/xsx/TCAM/9', '/xsx/TCAM/10', '/xsx/TCAM/11', '/xsx/TCAM/12', '/xsx/TCAM/13', '/xsx/TCAM/14', '/xsx/TCAM/15', '/xsx/TCAM/16', '/xsx/TCAM/17', '/xsx/TCAM/18'], dtype='object', name='tangential_cam_channel')) - tangential_cam_geometry_channelPandasIndex
PandasIndex(Index(['tangential_cam_1', 'tangential_cam_2', 'tangential_cam_3', 'tangential_cam_4', 'tangential_cam_5', 'tangential_cam_6', 'tangential_cam_7', 'tangential_cam_8', 'tangential_cam_9', 'tangential_cam_10', 'tangential_cam_11', 'tangential_cam_12', 'tangential_cam_13', 'tangential_cam_14', 'tangential_cam_15', 'tangential_cam_16', 'tangential_cam_17', 'tangential_cam_18'], dtype='object', name='tangential_cam_geometry_channel')) - third_horizontal_cam_geometry_channelPandasIndex
PandasIndex(Index(['third_horizontal_cam_1', 'third_horizontal_cam_2', 'third_horizontal_cam_3', 'third_horizontal_cam_4', 'third_horizontal_cam_5', 'third_horizontal_cam_6', 'third_horizontal_cam_7', 'third_horizontal_cam_8', 'third_horizontal_cam_9', 'third_horizontal_cam_10', 'third_horizontal_cam_11', 'third_horizontal_cam_12', 'third_horizontal_cam_13', 'third_horizontal_cam_14', 'third_horizontal_cam_15', 'third_horizontal_cam_16', 'third_horizontal_cam_17', 'third_horizontal_cam_18'], dtype='object', name='third_horizontal_cam_geometry_channel')) - timePandasIndex
PandasIndex(Index([ -0.06120026111602783, -0.06118026111602783, -0.061160261116027834, -0.061140261116027834, -0.061120261116027835, -0.061100261116027836, -0.06108026111602784, -0.06106026111602784, -0.06104026111602784, -0.06102026111602784, ... 0.6650197388839426, 0.6650397388839426, 0.6650597388839425, 0.6650797388839426, 0.6650997388839426, 0.6651197388839426, 0.6651397388839426, 0.6651597388839425, 0.6651797388839426, 0.6651997388839426], dtype='float64', name='time', length=36321))
- name :
- soft_x_rays
- description :
- Soft X-rays tomography diagnostic
- imas :
- soft_x_rays
- license_name :
- Creative Commons 4.0 BY-SA
- license_url :
- https://creativecommons.org/licenses/by-sa/4.0/
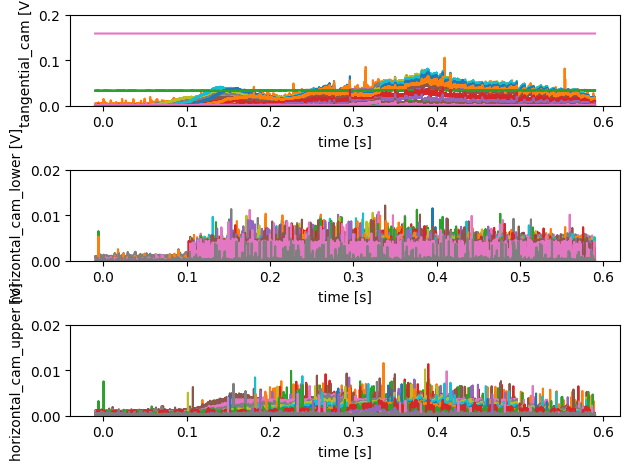
Thomson Profiles#
Thomson scattering measurements in a tokamak provide information about the plasma’s electron temperature and density profiles. The diagnostic analyses the scattering of laser light off free electrons in the plasma from a number of radial channels.
Below we plot the following profiles measured by the Thomson diagnostic
\(T_e\) - Electron temperature
\(N_e\) - Electron density
\(P_e\) - Electron pressure
profiles = xr.open_zarr(store, group='thomson_scattering')
profiles
fig, axes = plt.subplots(3, 1)
axes = axes.flatten()
profiles.t_e.plot(x='time', y='major_radius', ax=axes[0])
profiles.n_e.plot(x='time', y='major_radius', ax=axes[1])
profiles.p_e.plot(x='time', y='major_radius', ax=axes[2])
plt.tight_layout()
profiles
<xarray.Dataset> Size: 425kB
Dimensions: (major_radius: 120, time: 146)
Coordinates:
* major_radius (major_radius) float64 960B 0.3 0.31 0.32 ... 1.47 1.48 1.49
* time (time) float64 1kB -0.0612 -0.0562 -0.0512 ... 0.6588 0.6638
Data variables:
n_e (major_radius, time) float64 140kB ...
n_e_core (time) float64 1kB ...
p_e (major_radius, time) float64 140kB ...
t_e (major_radius, time) float64 140kB ...
t_e_core (time) float64 1kB ...
Attributes:
name: thomson_scattering
description: Thomson scattering diagnostic
imas: thomson_scattering
license_name: Creative Commons 4.0 BY-SA
license_url: https://creativecommons.org/licenses/by-sa/4.0/- major_radius: 120
- time: 146
- major_radius(major_radius)float640.3 0.31 0.32 ... 1.47 1.48 1.49
- description :
- label :
- major radius
- imas :
- thomson_scattering.channel[:].position.r
- units :
- m
array([0.3 , 0.31, 0.32, 0.33, 0.34, 0.35, 0.36, 0.37, 0.38, 0.39, 0.4 , 0.41, 0.42, 0.43, 0.44, 0.45, 0.46, 0.47, 0.48, 0.49, 0.5 , 0.51, 0.52, 0.53, 0.54, 0.55, 0.56, 0.57, 0.58, 0.59, 0.6 , 0.61, 0.62, 0.63, 0.64, 0.65, 0.66, 0.67, 0.68, 0.69, 0.7 , 0.71, 0.72, 0.73, 0.74, 0.75, 0.76, 0.77, 0.78, 0.79, 0.8 , 0.81, 0.82, 0.83, 0.84, 0.85, 0.86, 0.87, 0.88, 0.89, 0.9 , 0.91, 0.92, 0.93, 0.94, 0.95, 0.96, 0.97, 0.98, 0.99, 1. , 1.01, 1.02, 1.03, 1.04, 1.05, 1.06, 1.07, 1.08, 1.09, 1.1 , 1.11, 1.12, 1.13, 1.14, 1.15, 1.16, 1.17, 1.18, 1.19, 1.2 , 1.21, 1.22, 1.23, 1.24, 1.25, 1.26, 1.27, 1.28, 1.29, 1.3 , 1.31, 1.32, 1.33, 1.34, 1.35, 1.36, 1.37, 1.38, 1.39, 1.4 , 1.41, 1.42, 1.43, 1.44, 1.45, 1.46, 1.47, 1.48, 1.49]) - time(time)float64-0.0612 -0.0562 ... 0.6588 0.6638
- units :
- s
array([-0.0612, -0.0562, -0.0512, -0.0462, -0.0412, -0.0362, -0.0312, -0.0262, -0.0212, -0.0162, -0.0112, -0.0062, -0.0012, 0.0038, 0.0088, 0.0138, 0.0188, 0.0238, 0.0288, 0.0338, 0.0388, 0.0438, 0.0488, 0.0538, 0.0588, 0.0638, 0.0688, 0.0738, 0.0788, 0.0838, 0.0888, 0.0938, 0.0988, 0.1038, 0.1088, 0.1138, 0.1188, 0.1238, 0.1288, 0.1338, 0.1388, 0.1438, 0.1488, 0.1538, 0.1588, 0.1638, 0.1688, 0.1738, 0.1788, 0.1838, 0.1888, 0.1938, 0.1988, 0.2038, 0.2088, 0.2138, 0.2188, 0.2238, 0.2288, 0.2338, 0.2388, 0.2438, 0.2488, 0.2538, 0.2588, 0.2638, 0.2688, 0.2738, 0.2788, 0.2838, 0.2888, 0.2938, 0.2988, 0.3038, 0.3088, 0.3138, 0.3188, 0.3238, 0.3288, 0.3338, 0.3388, 0.3438, 0.3488, 0.3538, 0.3588, 0.3638, 0.3688, 0.3738, 0.3788, 0.3838, 0.3888, 0.3938, 0.3988, 0.4038, 0.4088, 0.4138, 0.4188, 0.4238, 0.4288, 0.4338, 0.4388, 0.4438, 0.4488, 0.4538, 0.4588, 0.4638, 0.4688, 0.4738, 0.4788, 0.4838, 0.4888, 0.4938, 0.4988, 0.5038, 0.5088, 0.5138, 0.5188, 0.5238, 0.5288, 0.5338, 0.5388, 0.5438, 0.5488, 0.5538, 0.5588, 0.5638, 0.5688, 0.5738, 0.5788, 0.5838, 0.5888, 0.5938, 0.5988, 0.6038, 0.6088, 0.6138, 0.6188, 0.6238, 0.6288, 0.6338, 0.6388, 0.6438, 0.6488, 0.6538, 0.6588, 0.6638])
- n_e(major_radius, time)float64...
- data_block_size :
- 268520
- description :
- Electron density
- label :
- electron density
- name :
- n_e
- uda_name :
- AYC_NE
- imas :
- thomson_scattering.channel[:].n_e
- units :
- 1 / m ** 3
[17520 values with dtype=float64]
- n_e_core(time)float64...
- data_block_size :
- 116168
- description :
- label :
- core density
- name :
- n_e_core
- uda_name :
- AYC_NE_CORE
- units :
- 1 / m ** 3
[146 values with dtype=float64]
- p_e(major_radius, time)float64...
- data_block_size :
- 268520
- description :
- label :
- electron pressure
- name :
- p_e
- uda_name :
- AYC_PE
- units :
- u * a
[17520 values with dtype=float64]
- t_e(major_radius, time)float64...
- data_block_size :
- 268520
- description :
- label :
- electron temperature
- name :
- t_e
- uda_name :
- AYC_TE
- imas :
- thomson_scattering.channel[:].t_e
- units :
- eV
[17520 values with dtype=float64]
- t_e_core(time)float64...
- data_block_size :
- 116168
- description :
- label :
- core temperature
- name :
- t_e_core
- uda_name :
- AYC_TE_CORE
- units :
- eV
[146 values with dtype=float64]
- major_radiusPandasIndex
PandasIndex(Index([ 0.3, 0.31, 0.32, 0.33, 0.34, 0.35000000000000003, 0.36000000000000004, 0.37000000000000005, 0.38000000000000006, 0.39000000000000007, ... 1.400000000000001, 1.410000000000001, 1.420000000000001, 1.430000000000001, 1.440000000000001, 1.450000000000001, 1.460000000000001, 1.470000000000001, 1.480000000000001, 1.490000000000001], dtype='float64', name='major_radius', length=120)) - timePandasIndex
PandasIndex(Index([ -0.06120026111602783, -0.056200261116027835, -0.05120026111602784, -0.04620026111602784, -0.04120026111602784, -0.036200261116027845, -0.031200261116027847, -0.02620026111602785, -0.021200261116027852, -0.016200261116027855, ... 0.6187997388839719, 0.6237997388839718, 0.6287997388839718, 0.6337997388839718, 0.6387997388839718, 0.6437997388839718, 0.6487997388839718, 0.6537997388839718, 0.6587997388839718, 0.6637997388839718], dtype='float64', name='time', length=146))
- name :
- thomson_scattering
- description :
- Thomson scattering diagnostic
- imas :
- thomson_scattering
- license_name :
- Creative Commons 4.0 BY-SA
- license_url :
- https://creativecommons.org/licenses/by-sa/4.0/
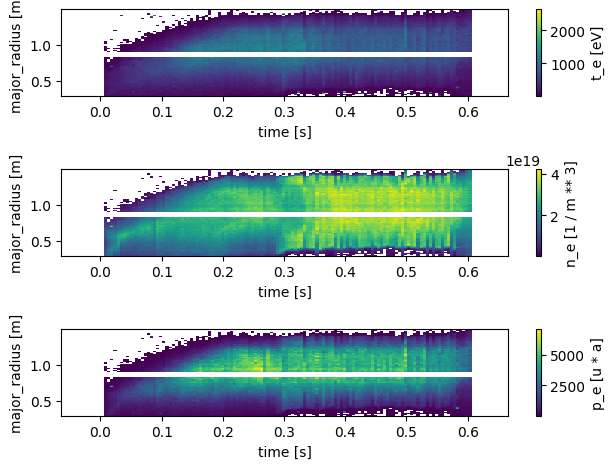
fig, axes = plt.subplots(2, 1)
profiles['t_e_core'].plot(x='time', ax=axes[0])
profiles['n_e_core'].plot(x='time', ax=axes[1])
for ax in axes:
ax.grid('on', alpha=0.5)
plt.tight_layout()
profiles
<xarray.Dataset> Size: 425kB
Dimensions: (major_radius: 120, time: 146)
Coordinates:
* major_radius (major_radius) float64 960B 0.3 0.31 0.32 ... 1.47 1.48 1.49
* time (time) float64 1kB -0.0612 -0.0562 -0.0512 ... 0.6588 0.6638
Data variables:
n_e (major_radius, time) float64 140kB ...
n_e_core (time) float64 1kB ...
p_e (major_radius, time) float64 140kB ...
t_e (major_radius, time) float64 140kB ...
t_e_core (time) float64 1kB ...
Attributes:
name: thomson_scattering
description: Thomson scattering diagnostic
imas: thomson_scattering
license_name: Creative Commons 4.0 BY-SA
license_url: https://creativecommons.org/licenses/by-sa/4.0/- major_radius: 120
- time: 146
- major_radius(major_radius)float640.3 0.31 0.32 ... 1.47 1.48 1.49
- description :
- label :
- major radius
- imas :
- thomson_scattering.channel[:].position.r
- units :
- m
array([0.3 , 0.31, 0.32, 0.33, 0.34, 0.35, 0.36, 0.37, 0.38, 0.39, 0.4 , 0.41, 0.42, 0.43, 0.44, 0.45, 0.46, 0.47, 0.48, 0.49, 0.5 , 0.51, 0.52, 0.53, 0.54, 0.55, 0.56, 0.57, 0.58, 0.59, 0.6 , 0.61, 0.62, 0.63, 0.64, 0.65, 0.66, 0.67, 0.68, 0.69, 0.7 , 0.71, 0.72, 0.73, 0.74, 0.75, 0.76, 0.77, 0.78, 0.79, 0.8 , 0.81, 0.82, 0.83, 0.84, 0.85, 0.86, 0.87, 0.88, 0.89, 0.9 , 0.91, 0.92, 0.93, 0.94, 0.95, 0.96, 0.97, 0.98, 0.99, 1. , 1.01, 1.02, 1.03, 1.04, 1.05, 1.06, 1.07, 1.08, 1.09, 1.1 , 1.11, 1.12, 1.13, 1.14, 1.15, 1.16, 1.17, 1.18, 1.19, 1.2 , 1.21, 1.22, 1.23, 1.24, 1.25, 1.26, 1.27, 1.28, 1.29, 1.3 , 1.31, 1.32, 1.33, 1.34, 1.35, 1.36, 1.37, 1.38, 1.39, 1.4 , 1.41, 1.42, 1.43, 1.44, 1.45, 1.46, 1.47, 1.48, 1.49]) - time(time)float64-0.0612 -0.0562 ... 0.6588 0.6638
- units :
- s
array([-0.0612, -0.0562, -0.0512, -0.0462, -0.0412, -0.0362, -0.0312, -0.0262, -0.0212, -0.0162, -0.0112, -0.0062, -0.0012, 0.0038, 0.0088, 0.0138, 0.0188, 0.0238, 0.0288, 0.0338, 0.0388, 0.0438, 0.0488, 0.0538, 0.0588, 0.0638, 0.0688, 0.0738, 0.0788, 0.0838, 0.0888, 0.0938, 0.0988, 0.1038, 0.1088, 0.1138, 0.1188, 0.1238, 0.1288, 0.1338, 0.1388, 0.1438, 0.1488, 0.1538, 0.1588, 0.1638, 0.1688, 0.1738, 0.1788, 0.1838, 0.1888, 0.1938, 0.1988, 0.2038, 0.2088, 0.2138, 0.2188, 0.2238, 0.2288, 0.2338, 0.2388, 0.2438, 0.2488, 0.2538, 0.2588, 0.2638, 0.2688, 0.2738, 0.2788, 0.2838, 0.2888, 0.2938, 0.2988, 0.3038, 0.3088, 0.3138, 0.3188, 0.3238, 0.3288, 0.3338, 0.3388, 0.3438, 0.3488, 0.3538, 0.3588, 0.3638, 0.3688, 0.3738, 0.3788, 0.3838, 0.3888, 0.3938, 0.3988, 0.4038, 0.4088, 0.4138, 0.4188, 0.4238, 0.4288, 0.4338, 0.4388, 0.4438, 0.4488, 0.4538, 0.4588, 0.4638, 0.4688, 0.4738, 0.4788, 0.4838, 0.4888, 0.4938, 0.4988, 0.5038, 0.5088, 0.5138, 0.5188, 0.5238, 0.5288, 0.5338, 0.5388, 0.5438, 0.5488, 0.5538, 0.5588, 0.5638, 0.5688, 0.5738, 0.5788, 0.5838, 0.5888, 0.5938, 0.5988, 0.6038, 0.6088, 0.6138, 0.6188, 0.6238, 0.6288, 0.6338, 0.6388, 0.6438, 0.6488, 0.6538, 0.6588, 0.6638])
- n_e(major_radius, time)float64...
- data_block_size :
- 268520
- description :
- Electron density
- label :
- electron density
- name :
- n_e
- uda_name :
- AYC_NE
- imas :
- thomson_scattering.channel[:].n_e
- units :
- 1 / m ** 3
[17520 values with dtype=float64]
- n_e_core(time)float64...
- data_block_size :
- 116168
- description :
- label :
- core density
- name :
- n_e_core
- uda_name :
- AYC_NE_CORE
- units :
- 1 / m ** 3
[146 values with dtype=float64]
- p_e(major_radius, time)float64...
- data_block_size :
- 268520
- description :
- label :
- electron pressure
- name :
- p_e
- uda_name :
- AYC_PE
- units :
- u * a
[17520 values with dtype=float64]
- t_e(major_radius, time)float64...
- data_block_size :
- 268520
- description :
- label :
- electron temperature
- name :
- t_e
- uda_name :
- AYC_TE
- imas :
- thomson_scattering.channel[:].t_e
- units :
- eV
[17520 values with dtype=float64]
- t_e_core(time)float64...
- data_block_size :
- 116168
- description :
- label :
- core temperature
- name :
- t_e_core
- uda_name :
- AYC_TE_CORE
- units :
- eV
[146 values with dtype=float64]
- major_radiusPandasIndex
PandasIndex(Index([ 0.3, 0.31, 0.32, 0.33, 0.34, 0.35000000000000003, 0.36000000000000004, 0.37000000000000005, 0.38000000000000006, 0.39000000000000007, ... 1.400000000000001, 1.410000000000001, 1.420000000000001, 1.430000000000001, 1.440000000000001, 1.450000000000001, 1.460000000000001, 1.470000000000001, 1.480000000000001, 1.490000000000001], dtype='float64', name='major_radius', length=120)) - timePandasIndex
PandasIndex(Index([ -0.06120026111602783, -0.056200261116027835, -0.05120026111602784, -0.04620026111602784, -0.04120026111602784, -0.036200261116027845, -0.031200261116027847, -0.02620026111602785, -0.021200261116027852, -0.016200261116027855, ... 0.6187997388839719, 0.6237997388839718, 0.6287997388839718, 0.6337997388839718, 0.6387997388839718, 0.6437997388839718, 0.6487997388839718, 0.6537997388839718, 0.6587997388839718, 0.6637997388839718], dtype='float64', name='time', length=146))
- name :
- thomson_scattering
- description :
- Thomson scattering diagnostic
- imas :
- thomson_scattering
- license_name :
- Creative Commons 4.0 BY-SA
- license_url :
- https://creativecommons.org/licenses/by-sa/4.0/
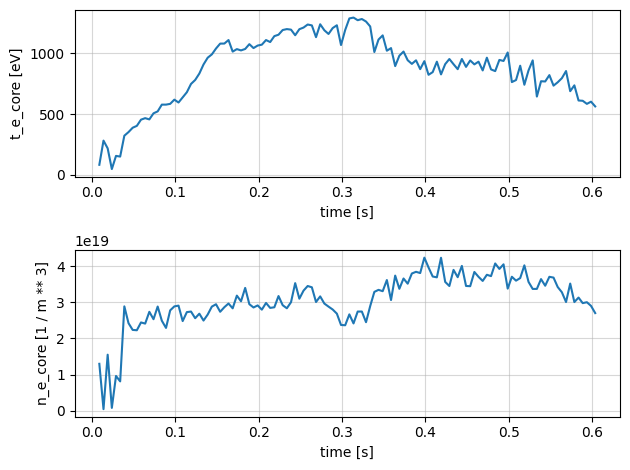
CXRS Profiles#
Charge Exchange Recombination Spectroscopy (CXRS) measurements provide information about ion temperature and plasma rotation. This diagnostic analyses the light emitted from charge exchange reactions between injected neutral beams and plasma ions.
Below we plot the following profiles measured by the CXRS diagnostic
\(T_i\) - Ion temperature
\(V_i\) - Ion velocity
profiles = xr.open_zarr(store, group='charge_exchange')
fig, axes = plt.subplots(2, 1)
profiles['t_i'].plot(x='time', y='major_radius', ax=axes[0], vmax=1000)
profiles['v_i'].plot(x='time', y='major_radius', ax=axes[1], vmax=1000)
plt.tight_layout()
profiles
<xarray.Dataset> Size: 376kB
Dimensions: (major_radius: 160, time: 146)
Coordinates:
* major_radius (major_radius) float64 1kB 0.0 0.01 0.02 ... 1.57 1.58 1.59
* time (time) float64 1kB -0.0612 -0.0562 -0.0512 ... 0.6588 0.6638
Data variables:
t_i (major_radius, time) float64 187kB ...
v_i (major_radius, time) float64 187kB ...
Attributes:
name: charge_exchange
description:
imas: charge_exchange
license_name: Creative Commons 4.0 BY-SA
license_url: https://creativecommons.org/licenses/by-sa/4.0/- major_radius: 160
- time: 146
- major_radius(major_radius)float640.0 0.01 0.02 ... 1.57 1.58 1.59
- units :
- m
array([0. , 0.01, 0.02, 0.03, 0.04, 0.05, 0.06, 0.07, 0.08, 0.09, 0.1 , 0.11, 0.12, 0.13, 0.14, 0.15, 0.16, 0.17, 0.18, 0.19, 0.2 , 0.21, 0.22, 0.23, 0.24, 0.25, 0.26, 0.27, 0.28, 0.29, 0.3 , 0.31, 0.32, 0.33, 0.34, 0.35, 0.36, 0.37, 0.38, 0.39, 0.4 , 0.41, 0.42, 0.43, 0.44, 0.45, 0.46, 0.47, 0.48, 0.49, 0.5 , 0.51, 0.52, 0.53, 0.54, 0.55, 0.56, 0.57, 0.58, 0.59, 0.6 , 0.61, 0.62, 0.63, 0.64, 0.65, 0.66, 0.67, 0.68, 0.69, 0.7 , 0.71, 0.72, 0.73, 0.74, 0.75, 0.76, 0.77, 0.78, 0.79, 0.8 , 0.81, 0.82, 0.83, 0.84, 0.85, 0.86, 0.87, 0.88, 0.89, 0.9 , 0.91, 0.92, 0.93, 0.94, 0.95, 0.96, 0.97, 0.98, 0.99, 1. , 1.01, 1.02, 1.03, 1.04, 1.05, 1.06, 1.07, 1.08, 1.09, 1.1 , 1.11, 1.12, 1.13, 1.14, 1.15, 1.16, 1.17, 1.18, 1.19, 1.2 , 1.21, 1.22, 1.23, 1.24, 1.25, 1.26, 1.27, 1.28, 1.29, 1.3 , 1.31, 1.32, 1.33, 1.34, 1.35, 1.36, 1.37, 1.38, 1.39, 1.4 , 1.41, 1.42, 1.43, 1.44, 1.45, 1.46, 1.47, 1.48, 1.49, 1.5 , 1.51, 1.52, 1.53, 1.54, 1.55, 1.56, 1.57, 1.58, 1.59]) - time(time)float64-0.0612 -0.0562 ... 0.6588 0.6638
- units :
- s
array([-0.0612, -0.0562, -0.0512, -0.0462, -0.0412, -0.0362, -0.0312, -0.0262, -0.0212, -0.0162, -0.0112, -0.0062, -0.0012, 0.0038, 0.0088, 0.0138, 0.0188, 0.0238, 0.0288, 0.0338, 0.0388, 0.0438, 0.0488, 0.0538, 0.0588, 0.0638, 0.0688, 0.0738, 0.0788, 0.0838, 0.0888, 0.0938, 0.0988, 0.1038, 0.1088, 0.1138, 0.1188, 0.1238, 0.1288, 0.1338, 0.1388, 0.1438, 0.1488, 0.1538, 0.1588, 0.1638, 0.1688, 0.1738, 0.1788, 0.1838, 0.1888, 0.1938, 0.1988, 0.2038, 0.2088, 0.2138, 0.2188, 0.2238, 0.2288, 0.2338, 0.2388, 0.2438, 0.2488, 0.2538, 0.2588, 0.2638, 0.2688, 0.2738, 0.2788, 0.2838, 0.2888, 0.2938, 0.2988, 0.3038, 0.3088, 0.3138, 0.3188, 0.3238, 0.3288, 0.3338, 0.3388, 0.3438, 0.3488, 0.3538, 0.3588, 0.3638, 0.3688, 0.3738, 0.3788, 0.3838, 0.3888, 0.3938, 0.3988, 0.4038, 0.4088, 0.4138, 0.4188, 0.4238, 0.4288, 0.4338, 0.4388, 0.4438, 0.4488, 0.4538, 0.4588, 0.4638, 0.4688, 0.4738, 0.4788, 0.4838, 0.4888, 0.4938, 0.4988, 0.5038, 0.5088, 0.5138, 0.5188, 0.5238, 0.5288, 0.5338, 0.5388, 0.5438, 0.5488, 0.5538, 0.5588, 0.5638, 0.5688, 0.5738, 0.5788, 0.5838, 0.5888, 0.5938, 0.5988, 0.6038, 0.6088, 0.6138, 0.6188, 0.6238, 0.6288, 0.6338, 0.6388, 0.6438, 0.6488, 0.6538, 0.6588, 0.6638])
- t_i(major_radius, time)float64...
- data_block_size :
- 183956
- description :
- Ion temperature at the channel measurement point
- label :
- Carbon temperature
- name :
- t_i
- uda_name :
- ACT_SS_TEMPERATURE
- imas :
- charge_exchange.channel[:].ion[:].t_i
- units :
- eV
[23360 values with dtype=float64]
- v_i(major_radius, time)float64...
- data_block_size :
- 183956
- description :
- label :
- Carbon Velocity
- name :
- v_i
- uda_name :
- ACT_SS_VELOCITY
- units :
- m / s
[23360 values with dtype=float64]
- major_radiusPandasIndex
PandasIndex(Index([ 0.0, 0.01, 0.02, 0.03, 0.04, 0.05, 0.06, 0.07, 0.08, 0.09, ... 1.5, 1.51, 1.52, 1.53, 1.54, 1.55, 1.56, 1.57, 1.58, 1.59], dtype='float64', name='major_radius', length=160)) - timePandasIndex
PandasIndex(Index([ -0.06120026111602783, -0.056200261116027835, -0.05120026111602784, -0.04620026111602784, -0.04120026111602784, -0.036200261116027845, -0.031200261116027847, -0.02620026111602785, -0.021200261116027852, -0.016200261116027855, ... 0.6187997388839719, 0.6237997388839718, 0.6287997388839718, 0.6337997388839718, 0.6387997388839718, 0.6437997388839718, 0.6487997388839718, 0.6537997388839718, 0.6587997388839718, 0.6637997388839718], dtype='float64', name='time', length=146))
- name :
- charge_exchange
- description :
- imas :
- charge_exchange
- license_name :
- Creative Commons 4.0 BY-SA
- license_url :
- https://creativecommons.org/licenses/by-sa/4.0/
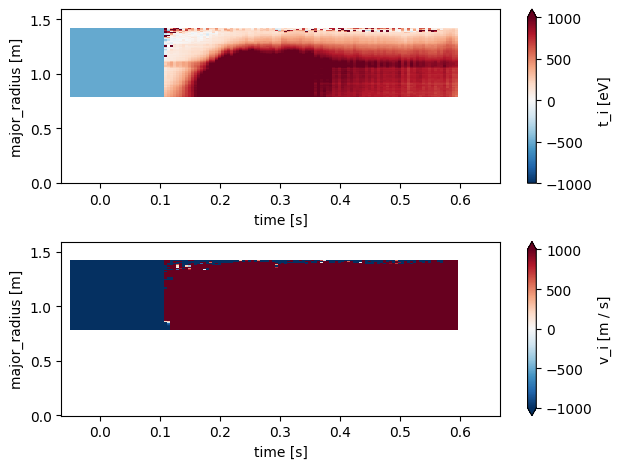
Equilibrium#
profiles = xr.open_zarr(store, group='equilibrium')
profile_1d = profiles[["beta_tor_normal", "wmhd", "li", "elongation", "triangularity_upper", "q95", "vloop_dynamic", "ip_rating", "lcfs_r", "lcfs_z"]]
plot_1d_profiles(profile_1d)
profiles
<xarray.Dataset> Size: 10MB
Dimensions: (time: 146, z: 65, major_radius: 65,
n_boundary_coords: 139, profile_r: 65,
n_x_points_r: 2, n_x_points_z: 2)
Coordinates:
* major_radius (major_radius) float64 520B 0.06 0.09 ... 1.95 1.98
* n_boundary_coords (n_boundary_coords) float32 556B 0.0 1.0 ... 138.0
* n_x_points_r (n_x_points_r) <U16 128B 'EFM_XPOINT1_R(C)' 'EFM_XPO...
* n_x_points_z (n_x_points_z) <U16 128B 'EFM_XPOINT1_Z(C)' 'EFM_XPO...
* profile_r (profile_r) float32 260B 0.0 0.01562 ... 0.9844 1.0
* time (time) float64 1kB -0.0612 -0.0562 ... 0.6588 0.6638
* z (z) float32 260B -2.0 -1.938 -1.875 ... 1.875 1.938 2.0
Data variables: (12/35)
beta_pol (time) float64 1kB ...
beta_tor (time) float64 1kB ...
beta_tor_normal (time) float64 1kB ...
bphi_rmag (time) float64 1kB ...
bvac_rmag (time) float64 1kB ...
da_rating (time) float64 1kB ...
... ...
triangularity_upper (time) float64 1kB ...
vloop_dynamic (time) float64 1kB ...
vloop_static (time) float64 1kB ...
wmhd (time) float64 1kB ...
x_point_r (n_x_points_r, time) float64 2kB ...
x_point_z (n_x_points_z, time) float64 2kB ...
Attributes:
name: equilibrium
description: Description of a 2D, axi-symmetric, tokamak equilibrium; r...
imas: equilibrium
license_name: Creative Commons 4.0 BY-SA
license_url: https://creativecommons.org/licenses/by-sa/4.0/- time: 146
- z: 65
- major_radius: 65
- n_boundary_coords: 139
- profile_r: 65
- n_x_points_r: 2
- n_x_points_z: 2
- major_radius(major_radius)float640.06 0.09 0.12 ... 1.92 1.95 1.98
- description :
- label :
- r-coordinates of computa
- imas :
- equilibrium.time_slice[:].coordinate_system.r
- units :
- m
array([0.06, 0.09, 0.12, 0.15, 0.18, 0.21, 0.24, 0.27, 0.3 , 0.33, 0.36, 0.39, 0.42, 0.45, 0.48, 0.51, 0.54, 0.57, 0.6 , 0.63, 0.66, 0.69, 0.72, 0.75, 0.78, 0.81, 0.84, 0.87, 0.9 , 0.93, 0.96, 0.99, 1.02, 1.05, 1.08, 1.11, 1.14, 1.17, 1.2 , 1.23, 1.26, 1.29, 1.32, 1.35, 1.38, 1.41, 1.44, 1.47, 1.5 , 1.53, 1.56, 1.59, 1.62, 1.65, 1.68, 1.71, 1.74, 1.77, 1.8 , 1.83, 1.86, 1.89, 1.92, 1.95, 1.98]) - n_boundary_coords(n_boundary_coords)float320.0 1.0 2.0 ... 136.0 137.0 138.0
- units :
array([ 0., 1., 2., 3., 4., 5., 6., 7., 8., 9., 10., 11., 12., 13., 14., 15., 16., 17., 18., 19., 20., 21., 22., 23., 24., 25., 26., 27., 28., 29., 30., 31., 32., 33., 34., 35., 36., 37., 38., 39., 40., 41., 42., 43., 44., 45., 46., 47., 48., 49., 50., 51., 52., 53., 54., 55., 56., 57., 58., 59., 60., 61., 62., 63., 64., 65., 66., 67., 68., 69., 70., 71., 72., 73., 74., 75., 76., 77., 78., 79., 80., 81., 82., 83., 84., 85., 86., 87., 88., 89., 90., 91., 92., 93., 94., 95., 96., 97., 98., 99., 100., 101., 102., 103., 104., 105., 106., 107., 108., 109., 110., 111., 112., 113., 114., 115., 116., 117., 118., 119., 120., 121., 122., 123., 124., 125., 126., 127., 128., 129., 130., 131., 132., 133., 134., 135., 136., 137., 138.], dtype=float32) - n_x_points_r(n_x_points_r)<U16'EFM_XPOINT1_R(C)' 'EFM_XPOINT2_...
array(['EFM_XPOINT1_R(C)', 'EFM_XPOINT2_R(C)'], dtype='<U16')
- n_x_points_z(n_x_points_z)<U16'EFM_XPOINT1_Z(C)' 'EFM_XPOINT2_...
array(['EFM_XPOINT1_Z(C)', 'EFM_XPOINT2_Z(C)'], dtype='<U16')
- profile_r(profile_r)float320.0 0.01562 0.03125 ... 0.9844 1.0
- units :
array([0. , 0.015625, 0.03125 , 0.046875, 0.0625 , 0.078125, 0.09375 , 0.109375, 0.125 , 0.140625, 0.15625 , 0.171875, 0.1875 , 0.203125, 0.21875 , 0.234375, 0.25 , 0.265625, 0.28125 , 0.296875, 0.3125 , 0.328125, 0.34375 , 0.359375, 0.375 , 0.390625, 0.40625 , 0.421875, 0.4375 , 0.453125, 0.46875 , 0.484375, 0.5 , 0.515625, 0.53125 , 0.546875, 0.5625 , 0.578125, 0.59375 , 0.609375, 0.625 , 0.640625, 0.65625 , 0.671875, 0.6875 , 0.703125, 0.71875 , 0.734375, 0.75 , 0.765625, 0.78125 , 0.796875, 0.8125 , 0.828125, 0.84375 , 0.859375, 0.875 , 0.890625, 0.90625 , 0.921875, 0.9375 , 0.953125, 0.96875 , 0.984375, 1. ], dtype=float32) - time(time)float64-0.0612 -0.0562 ... 0.6588 0.6638
- units :
- s
array([-0.0612, -0.0562, -0.0512, -0.0462, -0.0412, -0.0362, -0.0312, -0.0262, -0.0212, -0.0162, -0.0112, -0.0062, -0.0012, 0.0038, 0.0088, 0.0138, 0.0188, 0.0238, 0.0288, 0.0338, 0.0388, 0.0438, 0.0488, 0.0538, 0.0588, 0.0638, 0.0688, 0.0738, 0.0788, 0.0838, 0.0888, 0.0938, 0.0988, 0.1038, 0.1088, 0.1138, 0.1188, 0.1238, 0.1288, 0.1338, 0.1388, 0.1438, 0.1488, 0.1538, 0.1588, 0.1638, 0.1688, 0.1738, 0.1788, 0.1838, 0.1888, 0.1938, 0.1988, 0.2038, 0.2088, 0.2138, 0.2188, 0.2238, 0.2288, 0.2338, 0.2388, 0.2438, 0.2488, 0.2538, 0.2588, 0.2638, 0.2688, 0.2738, 0.2788, 0.2838, 0.2888, 0.2938, 0.2988, 0.3038, 0.3088, 0.3138, 0.3188, 0.3238, 0.3288, 0.3338, 0.3388, 0.3438, 0.3488, 0.3538, 0.3588, 0.3638, 0.3688, 0.3738, 0.3788, 0.3838, 0.3888, 0.3938, 0.3988, 0.4038, 0.4088, 0.4138, 0.4188, 0.4238, 0.4288, 0.4338, 0.4388, 0.4438, 0.4488, 0.4538, 0.4588, 0.4638, 0.4688, 0.4738, 0.4788, 0.4838, 0.4888, 0.4938, 0.4988, 0.5038, 0.5088, 0.5138, 0.5188, 0.5238, 0.5288, 0.5338, 0.5388, 0.5438, 0.5488, 0.5538, 0.5588, 0.5638, 0.5688, 0.5738, 0.5788, 0.5838, 0.5888, 0.5938, 0.5988, 0.6038, 0.6088, 0.6138, 0.6188, 0.6238, 0.6288, 0.6338, 0.6388, 0.6438, 0.6488, 0.6538, 0.6588, 0.6638]) - z(z)float32-2.0 -1.938 -1.875 ... 1.938 2.0
- description :
- label :
- z-coordinates of computa
- imas :
- equilibrium.time_slice[:].coordinate_system.z
- units :
- m
array([-2. , -1.9375, -1.875 , -1.8125, -1.75 , -1.6875, -1.625 , -1.5625, -1.5 , -1.4375, -1.375 , -1.3125, -1.25 , -1.1875, -1.125 , -1.0625, -1. , -0.9375, -0.875 , -0.8125, -0.75 , -0.6875, -0.625 , -0.5625, -0.5 , -0.4375, -0.375 , -0.3125, -0.25 , -0.1875, -0.125 , -0.0625, 0. , 0.0625, 0.125 , 0.1875, 0.25 , 0.3125, 0.375 , 0.4375, 0.5 , 0.5625, 0.625 , 0.6875, 0.75 , 0.8125, 0.875 , 0.9375, 1. , 1.0625, 1.125 , 1.1875, 1.25 , 1.3125, 1.375 , 1.4375, 1.5 , 1.5625, 1.625 , 1.6875, 1.75 , 1.8125, 1.875 , 1.9375, 2. ], dtype=float32)
- beta_pol(time)float64...
- data_block_size :
- 115724
- description :
- Poloidal beta. Defined as betap = 4 int(p dV) / [R_0 * mu_0 * Ip^2]
- label :
- Poloidal Beta
- name :
- beta_pol
- uda_name :
- EFM_BETAP
- imas :
- equilibrium.time_slice[:].global_quantities.beta_pol
- units :
[146 values with dtype=float64]
- beta_tor(time)float64...
- data_block_size :
- 115724
- description :
- Toroidal beta, defined as the volume-averaged total perpendicular pressure divided by (B0^2/(2*mu0)), i.e. beta_toroidal = 2 mu0 int(p dV) / V / B0^2
- label :
- Toroidal Beta
- name :
- beta_tor
- uda_name :
- EFM_BETAT
- imas :
- equilibrium.time_slice[:].global_quantities.beta_tor
- units :
[146 values with dtype=float64]
- beta_tor_normal(time)float64...
- data_block_size :
- 115724
- description :
- Normalized toroidal beta, defined as 100 * beta_tor * a[m] * B0 [T] / ip [MA]
- label :
- betat/(I/ a Bvac_geom)
- name :
- beta_tor_normal
- uda_name :
- EFM_BETAN
- imas :
- equilibrium.time_slice[:].global_quantities.beta_tor_normal
- units :
- T
[146 values with dtype=float64]
- bphi_rmag(time)float64...
- data_block_size :
- 115724
- description :
- label :
- Bphi at rmag
- name :
- bphi_rmag
- uda_name :
- EFM_BPHI_RMAG
- units :
- T
[146 values with dtype=float64]
- bvac_rmag(time)float64...
- data_block_size :
- 115724
- description :
- label :
- Bvac at rmag
- name :
- bvac_rmag
- uda_name :
- EFM_BVAC_RMAG
- units :
- T
[146 values with dtype=float64]
- da_rating(time)float64...
- data_block_size :
- 115724
- description :
- label :
- EFIT rating by D-alpha p
- name :
- da_rating
- uda_name :
- ESM_EFM_RATING_DA
- units :
[146 values with dtype=float64]
- elongation(time)float64...
- data_block_size :
- 115724
- description :
- Elongation of the plasma boundary
- label :
- Elongation
- name :
- elongation
- uda_name :
- EFM_ELONGATION
- imas :
- equilibrium.time_slice[:].boundary.elongation
- units :
[146 values with dtype=float64]
- elongation_axis(time)float64...
- data_block_size :
- 115724
- description :
- label :
- Elongation on Magnetic A
- name :
- elongation_axis
- uda_name :
- EFM_ELONGATION_AXIS
- units :
[146 values with dtype=float64]
- ip_rating(time)float64...
- data_block_size :
- 115724
- description :
- label :
- EFIT rating by I_p
- name :
- ip_rating
- uda_name :
- ESM_EFM_RATING_IP
- units :
[146 values with dtype=float64]
- j_phi(z, major_radius, time)float64...
- data_block_size :
- 3770164
- description :
- Toroidal plasma current density
- label :
- J(r,z)
- name :
- j_phi
- uda_name :
- EFM_PLASMA_CURR(R,Z)
- imas :
- equilibrium.time_slice[:].profiles_2d[:].j_phi
- units :
- A / m ** 2
[616850 values with dtype=float64]
- lcfs_r(n_boundary_coords, time)float64...
- data_block_size :
- 2373852
- description :
- label :
- r-coords of separatrix
- name :
- lcfs_r
- uda_name :
- EFM_LCFS(R)_(C)
- imas :
- equilibrium.time_slice[:].boundary.lcfs.r
- units :
- m
[20294 values with dtype=float64]
- lcfs_z(n_boundary_coords, time)float64...
- data_block_size :
- 2373852
- description :
- label :
- z-coords of separatrix
- name :
- lcfs_z
- uda_name :
- EFM_LCFS(Z)_(C)
- imas :
- equilibrium.time_slice[:].boundary.lcfs.z
- units :
- m
[20294 values with dtype=float64]
- li(time)float64...
- data_block_size :
- 115724
- description :
- Internal inductance
- label :
- Plasma Inductance
- name :
- li
- uda_name :
- EFM_LI
- imas :
- equilibrium.time_slice[:].global_quantities.li_3
- units :
[146 values with dtype=float64]
- magnetic_axis_r(time)float64...
- data_block_size :
- 115724
- description :
- Major radius of the magnetic axis
- label :
- Radius of Magnetic Axis
- name :
- magnetic_axis_r
- uda_name :
- EFM_MAGNETIC_AXIS_R
- imas :
- equilibrium.time_slice[:].global_quantities.magnetic_axis.r
- units :
- m
[146 values with dtype=float64]
- magnetic_axis_z(time)float64...
- data_block_size :
- 115724
- description :
- Height of the magnetic axis
- label :
- Height of Magnetic Axis
- name :
- magnetic_axis_z
- uda_name :
- EFM_MAGNETIC_AXIS_Z
- imas :
- equilibrium.time_slice[:].global_quantities.magnetic_axis.z
- units :
- m
[146 values with dtype=float64]
- minor_radius(time)float64...
- data_block_size :
- 115724
- description :
- Minor radius of the plasma boundary (defined as (Rmax-Rmin) / 2 of the boundary)
- label :
- Minor Radius
- name :
- minor_radius
- uda_name :
- EFM_MINOR_RADIUS
- imas :
- equilibrium.time_slice[:].boundary.minor_radius
- units :
- m
[146 values with dtype=float64]
- psi(z, major_radius, time)float64...
- data_block_size :
- 4175812
- description :
- Values of the poloidal flux at the grid in the poloidal plane. The poloidal flux is integral of magnetic field passing through a contour defined by the intersection of a flux surface passing through the point of interest and a Z=constant plane. If the integration surface is flat, the surface normal vector is in the increasing vertical coordinate direction, Z, namely upwards.
- label :
- psi(r,z)
- name :
- psi
- uda_name :
- EFM_PSI(R,Z)
- imas :
- equilibrium.time_slice[:].profiles_2d[:].psi
- units :
- Wb / rad
[616850 values with dtype=float64]
- q(profile_r, time)float64...
- data_block_size :
- 173472
- description :
- Safety factor (only positive when toroidal current and magnetic field are in same direction)
- label :
- q(r) at z=0.
- name :
- q
- uda_name :
- EFM_Q(R)
- imas :
- equilibrium.time_slice[:].profiles_1d.q
- units :
[9490 values with dtype=float64]
- q100(time)float64...
- data_block_size :
- 115724
- description :
- label :
- q at Psi_norm=100%
- name :
- q100
- uda_name :
- EFM_Q_100
- units :
[146 values with dtype=float64]
- q90(time)float64...
- data_block_size :
- 115724
- description :
- label :
- q at Psi_norm=100%
- name :
- q90
- uda_name :
- EFM_Q_100
- units :
[146 values with dtype=float64]
- q95(time)float64...
- data_block_size :
- 115724
- description :
- q at the 95% poloidal flux surface (only positive when toroidal current and magnetic field are in same direction)
- label :
- q at Psi_norm=95%
- name :
- q95
- uda_name :
- EFM_Q_95
- imas :
- equilibrium.time_slice[:].global_quantities.q_95
- units :
[146 values with dtype=float64]
- q_axis(time)float64...
- data_block_size :
- 115724
- description :
- q at the magnetic axis
- label :
- q on Magnetic Axis
- name :
- q_axis
- uda_name :
- EFM_Q_AXIS
- imas :
- equilibrium.time_slice[:].global_quantities.q_axis
- units :
[146 values with dtype=float64]
- rpsi100_in(time)float64...
- data_block_size :
- 115724
- description :
- label :
- i/b Rad of lcfs at Mid
- name :
- rpsi100_in
- uda_name :
- EFM_R(PSI100)_IN
- units :
- m
[146 values with dtype=float64]
- rpsi100_out(time)float64...
- data_block_size :
- 115724
- description :
- label :
- o/b Rad of lcfs at Mid
- name :
- rpsi100_out
- uda_name :
- EFM_R(PSI100)_OUT
- units :
- m
[146 values with dtype=float64]
- rpsi90_in(time)float64...
- data_block_size :
- 115724
- description :
- label :
- i/b Rad of 90%Psi at Mid
- name :
- rpsi90_in
- uda_name :
- EFM_R(PSI90)_IN
- units :
- m
[146 values with dtype=float64]
- rpsi90_out(time)float64...
- data_block_size :
- 115724
- description :
- label :
- o/b Rad of 90%Psi at Mid
- name :
- rpsi90_out
- uda_name :
- EFM_R(PSI90)_OUT
- units :
- m
[146 values with dtype=float64]
- rpsi95_in(time)float64...
- data_block_size :
- 115724
- description :
- label :
- i/b Rad of 95%Psi at Mid
- name :
- rpsi95_in
- uda_name :
- EFM_R(PSI95)_IN
- units :
- m
[146 values with dtype=float64]
- rpsi95_out(time)float64...
- data_block_size :
- 115724
- description :
- label :
- o/b Rad of 95%Psi at Mid
- name :
- rpsi95_out
- uda_name :
- EFM_R(PSI95)_OUT
- units :
- m
[146 values with dtype=float64]
- triangularity_lower(time)float64...
- data_block_size :
- 115724
- description :
- Lower triangularity w.r.t. magnetic axis
- label :
- Lower Triangularity
- name :
- triangularity_lower
- uda_name :
- EFM_TRIANG_LOWER
- imas :
- equilibrium.time_slice[:].profiles_1d.triangularity_lower
- units :
[146 values with dtype=float64]
- triangularity_upper(time)float64...
- data_block_size :
- 115724
- description :
- Upper triangularity w.r.t. magnetic axis
- label :
- Upper Triangularity
- name :
- triangularity_upper
- uda_name :
- EFM_TRIANG_UPPER
- imas :
- equilibrium.time_slice[:].profiles_1d.triangularity_upper
- units :
[146 values with dtype=float64]
- vloop_dynamic(time)float64...
- data_block_size :
- 115724
- description :
- label :
- dynamic V_loop
- name :
- vloop_dynamic
- uda_name :
- ESM_V_LOOP_DYNAMIC
- units :
- V
[146 values with dtype=float64]
- vloop_static(time)float64...
- data_block_size :
- 115724
- description :
- label :
- static V_loop
- name :
- vloop_static
- uda_name :
- ESM_V_LOOP_STATIC
- units :
- V
[146 values with dtype=float64]
- wmhd(time)float64...
- data_block_size :
- 115724
- description :
- label :
- Plasma Diamagnetic Energ
- name :
- wmhd
- uda_name :
- EFM_WPLASMD
- units :
- J
[146 values with dtype=float64]
- x_point_r(n_x_points_r, time)float64...
- data_block_size :
- 115724
- description :
- label :
- Radius of X-point 1
- name :
- x_point_r
- uda_name :
- EFM_XPOINT1_R(C)
- imas :
- equilibrium.time_slice[:].boundary.x_point[:].r
- units :
[292 values with dtype=float64]
- x_point_z(n_x_points_z, time)float64...
- data_block_size :
- 115724
- description :
- label :
- Height of X-point 1
- name :
- x_point_z
- uda_name :
- EFM_XPOINT1_Z(C)
- imas :
- equilibrium.time_slice[:].boundary.x_point[:].z
- units :
[292 values with dtype=float64]
- major_radiusPandasIndex
PandasIndex(Index([ 0.06, 0.09, 0.12, 0.15, 0.18, 0.21, 0.24, 0.27, 0.3, 0.33, 0.36, 0.38999999999999996, 0.42, 0.45, 0.48, 0.51, 0.54, 0.5700000000000001, 0.6000000000000001, 0.6299999999999999, 0.6599999999999999, 0.69, 0.72, 0.75, 0.78, 0.81, 0.8400000000000001, 0.8699999999999999, 0.8999999999999999, 0.9299999999999999, 0.96, 0.99, 1.02, 1.05, 1.08, 1.11, 1.1400000000000001, 1.17, 1.2, 1.23, 1.26, 1.29, 1.32, 1.35, 1.38, 1.41, 1.44, 1.47, 1.5, 1.53, 1.56, 1.59, 1.62, 1.65, 1.68, 1.71, 1.74, 1.77, 1.8, 1.83, 1.8599999999999999, 1.89, 1.92, 1.95, 1.98], dtype='float64', name='major_radius')) - n_boundary_coordsPandasIndex
PandasIndex(Index([ 0.0, 1.0, 2.0, 3.0, 4.0, 5.0, 6.0, 7.0, 8.0, 9.0, ... 129.0, 130.0, 131.0, 132.0, 133.0, 134.0, 135.0, 136.0, 137.0, 138.0], dtype='float32', name='n_boundary_coords', length=139)) - n_x_points_rPandasIndex
PandasIndex(Index(['EFM_XPOINT1_R(C)', 'EFM_XPOINT2_R(C)'], dtype='object', name='n_x_points_r'))
- n_x_points_zPandasIndex
PandasIndex(Index(['EFM_XPOINT1_Z(C)', 'EFM_XPOINT2_Z(C)'], dtype='object', name='n_x_points_z'))
- profile_rPandasIndex
PandasIndex(Index([ 0.0, 0.015625, 0.03125, 0.046875, 0.0625, 0.078125, 0.09375, 0.109375, 0.125, 0.140625, 0.15625, 0.171875, 0.1875, 0.203125, 0.21875, 0.234375, 0.25, 0.265625, 0.28125, 0.296875, 0.3125, 0.328125, 0.34375, 0.359375, 0.375, 0.390625, 0.40625, 0.421875, 0.4375, 0.453125, 0.46875, 0.484375, 0.5, 0.515625, 0.53125, 0.546875, 0.5625, 0.578125, 0.59375, 0.609375, 0.625, 0.640625, 0.65625, 0.671875, 0.6875, 0.703125, 0.71875, 0.734375, 0.75, 0.765625, 0.78125, 0.796875, 0.8125, 0.828125, 0.84375, 0.859375, 0.875, 0.890625, 0.90625, 0.921875, 0.9375, 0.953125, 0.96875, 0.984375, 1.0], dtype='float32', name='profile_r')) - timePandasIndex
PandasIndex(Index([ -0.06120026111602783, -0.056200261116027835, -0.05120026111602784, -0.04620026111602784, -0.04120026111602784, -0.036200261116027845, -0.031200261116027847, -0.02620026111602785, -0.021200261116027852, -0.016200261116027855, ... 0.6187997388839719, 0.6237997388839718, 0.6287997388839718, 0.6337997388839718, 0.6387997388839718, 0.6437997388839718, 0.6487997388839718, 0.6537997388839718, 0.6587997388839718, 0.6637997388839718], dtype='float64', name='time', length=146)) - zPandasIndex
PandasIndex(Index([ -2.0, -1.9375, -1.875, -1.8125, -1.75, -1.6875, -1.625, -1.5625, -1.5, -1.4375, -1.375, -1.3125, -1.25, -1.1875, -1.125, -1.0625, -1.0, -0.9375, -0.875, -0.8125, -0.75, -0.6875, -0.625, -0.5625, -0.5, -0.4375, -0.375, -0.3125, -0.25, -0.1875, -0.125, -0.0625, 0.0, 0.0625, 0.125, 0.1875, 0.25, 0.3125, 0.375, 0.4375, 0.5, 0.5625, 0.625, 0.6875, 0.75, 0.8125, 0.875, 0.9375, 1.0, 1.0625, 1.125, 1.1875, 1.25, 1.3125, 1.375, 1.4375, 1.5, 1.5625, 1.625, 1.6875, 1.75, 1.8125, 1.875, 1.9375, 2.0], dtype='float32', name='z'))
- name :
- equilibrium
- description :
- Description of a 2D, axi-symmetric, tokamak equilibrium; result of an equilibrium code.
- imas :
- equilibrium
- license_name :
- Creative Commons 4.0 BY-SA
- license_url :
- https://creativecommons.org/licenses/by-sa/4.0/
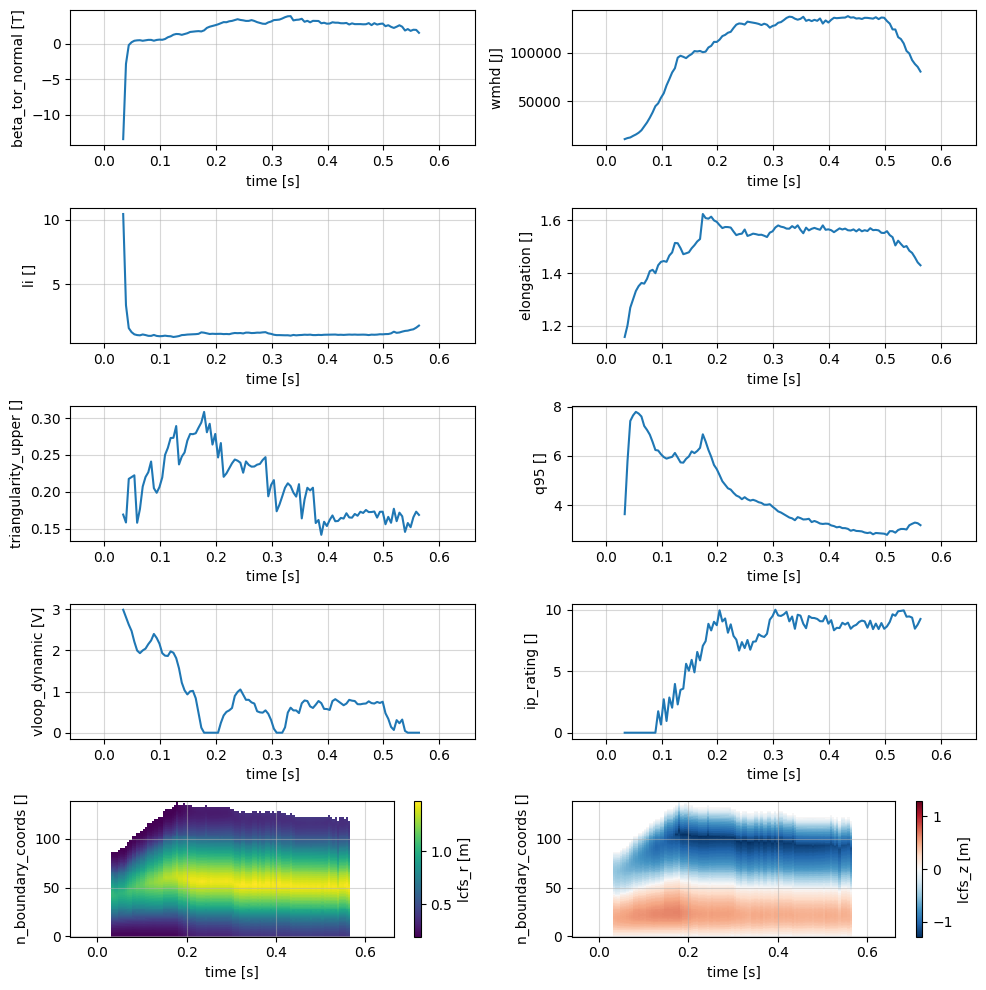
fig, axes = plt.subplots(1, 3, figsize=(10, 5))
profiles['j_phi'].isel(time=50).plot(ax=axes[0], x='major_radius')
profiles['psi'].isel(time=50).plot(ax=axes[1], x='major_radius')
profiles['q'].isel(time=50).plot(ax=axes[2])
plt.tight_layout()
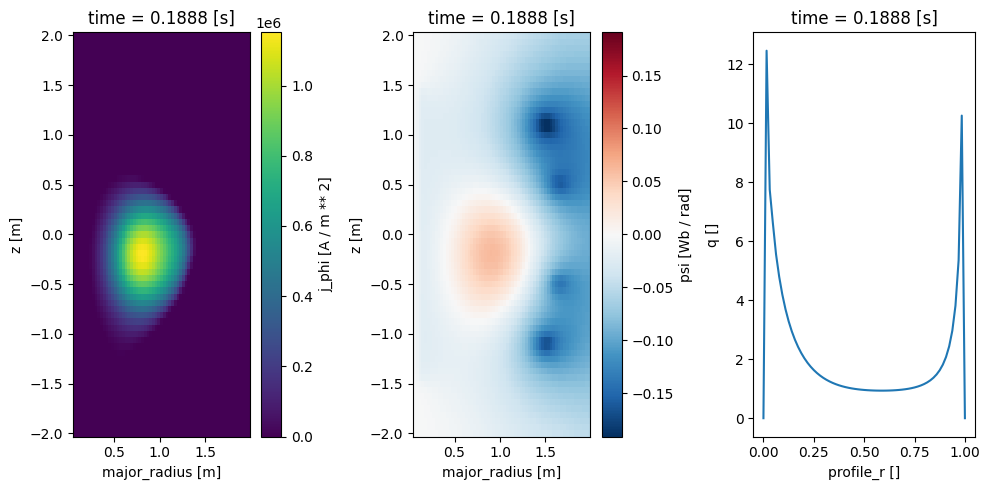
Gas Injection#
profiles = xr.open_zarr(store, group='gas_injection')
plot_1d_profiles(profiles[["inboard_total", "outboard_total", "pressure", "total_injected"]])
plt.tight_layout()
profiles
<xarray.Dataset> Size: 862kB
Dimensions: (time: 2906, target_valve_channel: 12,
valve_channel: 20)
Coordinates:
* target_valve_channel (target_valve_channel) <U14 672B '/xdc/gas/t/g1' .....
* time (time) float64 23kB -0.0612 -0.06095 ... 0.6648 0.665
* valve_channel (valve_channel) <U16 1kB '/xdc/gas/f/tc5a' ... '/xd...
Data variables:
inboard_total (time) float64 23kB ...
outboard_total (time) float64 23kB ...
pressure (time) float64 23kB ...
total_injected (time) float64 23kB ...
valve_target_voltage (target_valve_channel, time) float64 279kB ...
valve_voltage (valve_channel, time) float64 465kB ...
Attributes:
name: gas_injection
description: Gas injection by a system of pipes and valves.
imas: gas_injection
license_name: Creative Commons 4.0 BY-SA
license_url: https://creativecommons.org/licenses/by-sa/4.0/- time: 2906
- target_valve_channel: 12
- valve_channel: 20
- target_valve_channel(target_valve_channel)<U14'/xdc/gas/t/g1' ... '/xdc/gas/t/...
array(['/xdc/gas/t/g1', '/xdc/gas/t/g2', '/xdc/gas/t/g3', '/xdc/gas/t/g4', '/xdc/gas/t/g5', '/xdc/gas/t/g6', '/xdc/gas/t/g7', '/xdc/gas/t/g8', '/xdc/gas/t/g9', '/xdc/gas/t/g10', '/xdc/gas/t/g11', '/xdc/gas/t/g12'], dtype='<U14') - time(time)float64-0.0612 -0.06095 ... 0.6648 0.665
- units :
- s
array([-0.0612 , -0.06095, -0.0607 , ..., 0.66455, 0.6648 , 0.66505], shape=(2906,)) - valve_channel(valve_channel)<U16'/xdc/gas/f/tc5a' ... '/xdc/gas/...
array(['/xdc/gas/f/tc5a', '/xdc/gas/f/tc5b', '/xdc/gas/f/tc11', '/xdc/gas/f/bc5', '/xdc/gas/f/hl1', '/xdc/gas/f/hu6', '/xdc/gas/f/hu8', '/xdc/gas/f/hu11', '/xdc/gas/f/hl11', '/xdc/gas/f/bc11', '/xdc/gas/f/hecc', '/xdc/gas/f/heoa', '/xdc/gas/f/heob', '/xdc/gas/f/hm12a', '/xdc/gas/f/hm12b', '/xdc/gas/f/ibil', '/xdc/gas/f/ibfua', '/xdc/gas/f/ibfub', '/xdc/gas/f/ibfla', '/xdc/gas/f/ibflb'], dtype='<U16')
- inboard_total(time)float64...
- data_block_size :
- 210428
- description :
- label :
- Total inboard gas
- name :
- inboard_total
- uda_name :
- AGA_INBOARD_TOTAL
- units :
- cps
[2906 values with dtype=float64]
- outboard_total(time)float64...
- data_block_size :
- 210428
- description :
- label :
- Total outboard gas
- name :
- outboard_total
- uda_name :
- AGA_OUTBOARD_TOTAL
- units :
- cps
[2906 values with dtype=float64]
- pressure(time)float64...
- data_block_size :
- 1638428
- description :
- label :
- Fast Ion Gauge molec.den
- units :
- D2/m3
- name :
- pressure
- uda_name :
- AGA_FIG
[2906 values with dtype=float64]
- total_injected(time)float64...
- data_block_size :
- 210428
- description :
- label :
- Integrated total gas
- name :
- total_injected
- uda_name :
- AGA_INTEG_GAS
- units :
- count
[2906 values with dtype=float64]
- valve_target_voltage(target_valve_channel, time)float64...
- data_block_size :
- 132028
- description :
- label :
- /xdc/gas/t/g1
- name :
- valve_target_voltage
- uda_name :
- /xdc/gas/t/g1
- imas :
- gas_injection.valve[:].target_voltage.data
- units :
- V
[34872 values with dtype=float64]
- valve_voltage(valve_channel, time)float64...
- data_block_size :
- 450428
- description :
- label :
- /xdc/gas/f/tc5a
- name :
- valve_voltage
- uda_name :
- /xdc/gas/f/tc5a
- imas :
- gas_injection.valve[:].voltage.data
- fuelling_groups :
- {"/xdc/gas/t/g1": ["/xdc/gas/f/tc5a", "/xdc/gas/f/tc5b", "/xdc/gas/f/tc11"], "/xdc/gas/t/g2": ["/xdc/gas/f/bc5"], "/xdc/gas/t/g3": ["/xdc/gas/f/hl1", "/xdc/gas/f/hu6", "/xdc/gas/f/hu8"], "/xdc/gas/t/g4": ["/xdc/gas/f/hu11"], "/xdc/gas/t/g11": ["/xdc/gas/f/ibfua", "/xdc/gas/f/ibfub"], "/xdc/gas/t/g12": ["/xdc/gas/f/ibfla", "/xdc/gas/f/ibflb"]}
- impurity_groups :
- {"/xdc/gas/t/g5": ["/xdc/gas/f/hl11"], "/xdc/gas/t/g6": ["/xdc/gas/f/bc11"], "/xdc/gas/t/g7": ["/xdc/gas/f/hecc"], "/xdc/gas/t/g8": ["/xdc/gas/f/heoa", "/xdc/gas/f/heob"], "/xdc/gas/t/g9": ["/xdc/gas/f/hm12a", "/xdc/gas/f/hm12b"], "/xdc/gas/t/g10": ["/xdc/gas/f/ibil"]}
- units :
- V
[58120 values with dtype=float64]
- target_valve_channelPandasIndex
PandasIndex(Index(['/xdc/gas/t/g1', '/xdc/gas/t/g2', '/xdc/gas/t/g3', '/xdc/gas/t/g4', '/xdc/gas/t/g5', '/xdc/gas/t/g6', '/xdc/gas/t/g7', '/xdc/gas/t/g8', '/xdc/gas/t/g9', '/xdc/gas/t/g10', '/xdc/gas/t/g11', '/xdc/gas/t/g12'], dtype='object', name='target_valve_channel')) - timePandasIndex
PandasIndex(Index([-0.06120026111602783, -0.06095026111602783, -0.06070026111602783, -0.06045026111602783, -0.06020026111602783, -0.05995026111602783, -0.05970026111602783, -0.05945026111602783, -0.05920026111602783, -0.05895026111602783, ... 0.6627997388839728, 0.6630497388839728, 0.6632997388839728, 0.6635497388839728, 0.6637997388839728, 0.6640497388839728, 0.6642997388839729, 0.6645497388839728, 0.6647997388839728, 0.6650497388839728], dtype='float64', name='time', length=2906)) - valve_channelPandasIndex
PandasIndex(Index(['/xdc/gas/f/tc5a', '/xdc/gas/f/tc5b', '/xdc/gas/f/tc11', '/xdc/gas/f/bc5', '/xdc/gas/f/hl1', '/xdc/gas/f/hu6', '/xdc/gas/f/hu8', '/xdc/gas/f/hu11', '/xdc/gas/f/hl11', '/xdc/gas/f/bc11', '/xdc/gas/f/hecc', '/xdc/gas/f/heoa', '/xdc/gas/f/heob', '/xdc/gas/f/hm12a', '/xdc/gas/f/hm12b', '/xdc/gas/f/ibil', '/xdc/gas/f/ibfua', '/xdc/gas/f/ibfub', '/xdc/gas/f/ibfla', '/xdc/gas/f/ibflb'], dtype='object', name='valve_channel'))
- name :
- gas_injection
- description :
- Gas injection by a system of pipes and valves.
- imas :
- gas_injection
- license_name :
- Creative Commons 4.0 BY-SA
- license_url :
- https://creativecommons.org/licenses/by-sa/4.0/